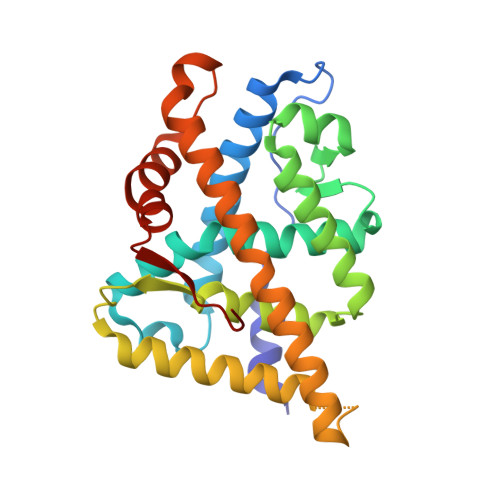
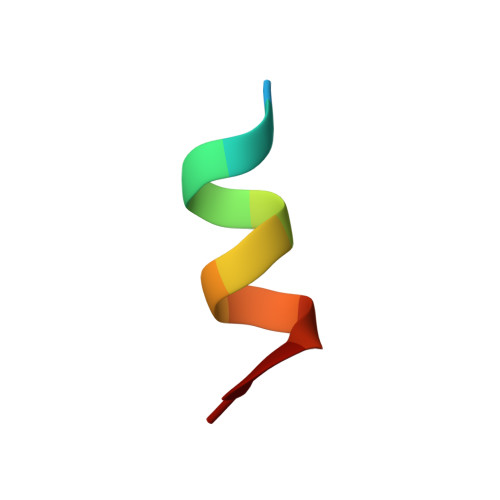Identification of a new androgen receptor (AR) co-regulator BUD31 and related peptides to suppress wild-type and mutated AR-mediated prostate cancer growth via peptide screening and X-ray structure analysis.
Hsu, C.L., Liu, J.S., Wu, P.L., Guan, H.H., Chen, Y.L., Lin, A.C., Ting, H.J., Pang, S.T., Yeh, S.D., Ma, W.L., Chen, C.J., Wu, W.G., Chang, C.(2014) Mol Oncol 8: 1575-1587
- PubMed: 25091737
- DOI: https://doi.org/10.1016/j.molonc.2014.06.009
- Primary Citation of Related Structures:
4OEA, 4OED, 4OEY, 4OEZ, 4OFR, 4OFU, 4OGH, 4OH5, 4OH6, 4OHA, 4OIL, 4OIU, 4OJ9, 4OJB, 4OK1, 4OKB, 4OKT, 4OKW, 4OKX, 4OLM - PubMed Abstract:
Treatment with individual anti-androgens is associated with the development of hot-spot mutations in the androgen receptor (AR). Here, we found that anti-androgens-mt-ARs have similar binary structure to the 5α-dihydrotestosterone-wt-AR. Phage display revealed that these ARs bound to similar peptides, including BUD31, containing an Fxx(F/H/L/W/Y)Y motif cluster with Tyr in the +5 position. Structural analyses of the AR-LBD-BUD31 complex revealed formation of an extra hydrogen bond between the Tyr+5 residue of the peptide and the AR. Functional studies showed that BUD31-related peptides suppressed AR transactivation, interrupted AR N-C interaction, and suppressed AR-mediated cell growth. Combination of peptide screening and X-ray structure analysis may serve as a new strategy for developing anti-ARs that simultaneously suppress both wt and mutated AR function.
Organizational Affiliation:
The George Whipple Lab for Cancer Research, Department of Pathology and Urology, University of Rochester Medical Center, Rochester, NY 14642, USA; Division of Hematology-Oncology, Department of Internal Medicine, Chang Gung Memorial Hospital, Chang Gung University, Taoyuan 333, Taiwan.