Pharmacological chaperones for GCase that switch conformation with pH enhance enzyme levels in Gaucher animal models.
Santana, A.G., Robinson, K., Vickers, C., Chen, H., Zhou, S., Boraston, A.B., Clarke, L.A., Withers, S.G.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Glucosylceramidase | 497 | Homo sapiens | Mutation(s): 0 EC: 3.2.1.45 (PDB Primary Data), 3.2.1.46 (UniProt), 2.4.1 (UniProt), 3.2.1 (UniProt) | 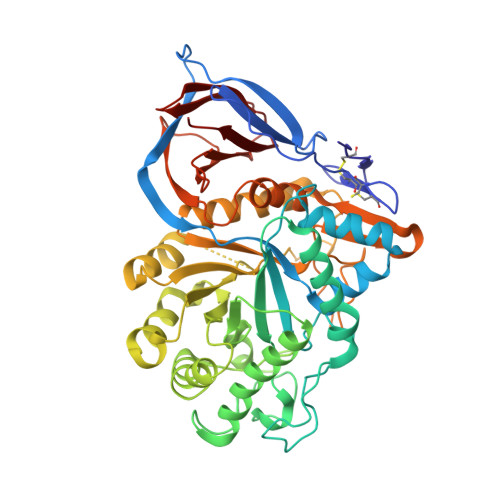 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P04062 (Homo sapiens) Explore P04062 Go to UniProtKB: P04062 | |||||
PHAROS: P04062 GTEx: ENSG00000177628 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P04062 | ||||
Glycosylation | |||||
| Glycosylation Sites: 3 | Go to GlyGen: P04062-1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | 3D Interactions |
| 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | C | 2 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G42666HT GlyCosmos: G42666HT GlyGen: G42666HT | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Query on NAG | W [auth B], X [auth B] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| P9V (Subject of Investigation/LOI) Query on P9V | PA [auth B], V [auth A] | (2R,3S,4S,5R)-2-[2-(methylsulfanyl)ethyl]piperidine-3,4,5-triol C8 H17 N O3 S ZTGUKWHBLOJEPU-NGJRWZKOSA-N |  | ||
| SO4 Query on SO4 | FA [auth B] GA [auth B] HA [auth B] IA [auth B] JA [auth B] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| EDO Query on EDO | AA [auth B] BA [auth B] CA [auth B] DA [auth B] EA [auth B] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 109.935 | α = 90 |
| b = 285.06 | β = 90 |
| c = 91.815 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data scaling |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Canadian Institutes of Health Research (CIHR) | Canada | -- |