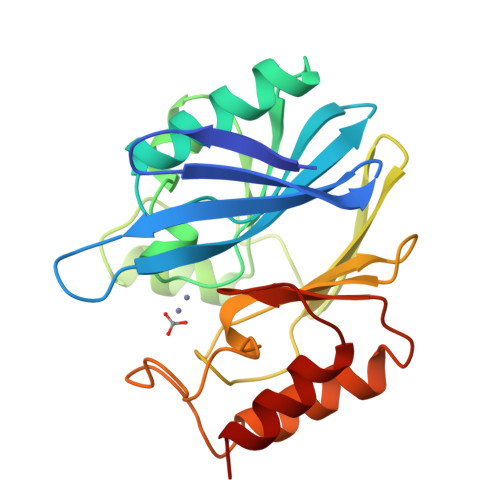
1.85 A resolution structure of the zinc (II) beta-lactamase from Bacillus cereus.
Carfi, A., Duee, E., Galleni, M., Frere, J.M., Dideberg, O.(1998) Acta Crystallogr D Biol Crystallogr 54: 313-323
- PubMed: 9761898
- DOI: https://doi.org/10.1107/s0907444997010627
- Primary Citation of Related Structures:
1BVT - PubMed Abstract:
Class B beta-lactamases are wide spectrum enzymes which require bivalent metal ions for activity. The structure of the class B zinc-ion-dependent beta-lactamase from Bacillus cereus (BCII) has been refined at 1.85 A resolution using data collected on cryocooled crystals (100 K). The enzyme from B. cereus has a molecular mass of 24 946 Da and is folded into a beta-sandwich structure with helices on the external faces. The active site is located in a groove running between the two beta-sheets [Carfi et al. (1995). EMBO J. 14, 4914-4921]. The 100 K high-resolution BCII structure shows one fully and one partially occupied zinc sites. The zinc ion in the fully occupied site (the catalytic zinc) is coordinated by three histidines and one water molecule. The second zinc ion is at 3.7 A from the first one and is coordinated by one histidine, one cysteine, one aspartate and one unknown molecule (most likely a carbonate ion). In the B. cereus zinc beta-lactamase the affinity for the second metal-ion is low at the pH of crystallization (Kd = 25 mM, 293 K; [Baldwin et al. (1978). Biochem. J. 175, 441-447] and the dissociation constant of the second zinc ion was thus apparently decreased at the cryogenic temperature. In addition, the structure of the apo enzyme was determined at 2.5 A resolution. The removal of the zinc ion by chelating agents causes small changes in the active-site environment.
Organizational Affiliation:
LCM, Institut de Biologie Structurale Jean-Pierre Ebel (CEA-CNRS), Grenoble, France.