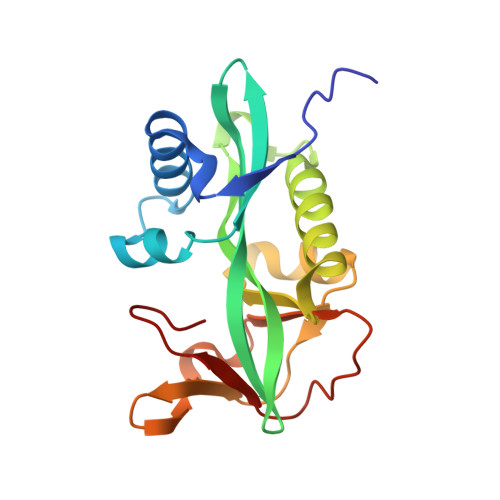
Aminoglycoside 2'-N-acetyltransferase from Mycobacterium tuberculosis in complex with coenzyme A and aminoglycoside substrates.
Vetting, M.W., Hegde, S.S., Javid-Majd, F., Blanchard, J.S., Roderick, S.L.(2002) Nat Struct Biol 9: 653-658
- PubMed: 12161746
- DOI: https://doi.org/10.1038/nsb830
- Primary Citation of Related Structures:
1M44, 1M4D, 1M4G, 1M4I - PubMed Abstract:
AAC(2')-Ic catalyzes the coenzyme A (CoA)-dependent acetylation of the 2' hydroxyl or amino group of a broad spectrum of aminoglycosides. The crystal structure of the AAC(2')-Ic from Mycobacterium tuberculosis has been determined in the apo enzyme form and in ternary complexes with CoA and either tobramycin, kanamycin A or ribostamycin, representing the first structures of an aminoglycoside acetyltransferase bound to a drug. The overall fold of AAC(2')-Ic places it in the GCN5-related N-acetyltransferase (GNAT) superfamily. Although the physiological function of AAC(2')-Ic is uncertain, a structural analysis of these high-affinity aminoglycoside complexes suggests that the enzyme may acetylate a key biosynthetic intermediate of mycothiol, the major reducing agent in mycobacteria, and participate in the regulation of cellular redox potential.
Organizational Affiliation:
Department of Biochemistry, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, New York 10461, USA.