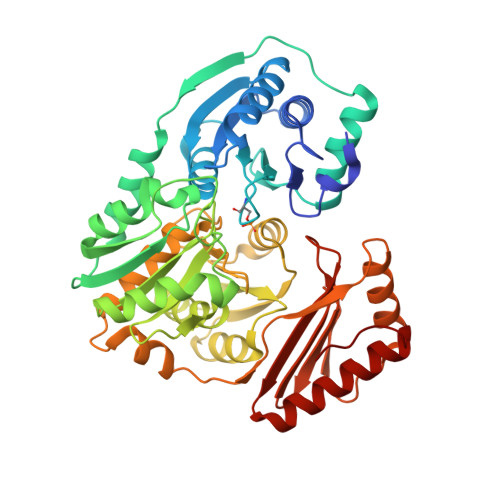
Structural basis of diverse substrate recognition by the enzyme PMM/PGM from P. aeruginosa.
Regni, C., Naught, L., Tipton, P.A., Beamer, L.J.(2004) Structure 12: 55-63
- PubMed: 14725765
- DOI: https://doi.org/10.1016/j.str.2003.11.015
- Primary Citation of Related Structures:
1P5D, 1P5G, 1PCJ, 1PCM - PubMed Abstract:
Enzyme-substrate complexes of phosphomannomutase/phosphoglucomutase (PMM/PGM) reveal the structural basis of the enzyme's ability to use four different substrates in catalysis. High-resolution structures with glucose 1-phosphate, glucose 6-phosphate, mannose 1-phosphate, and mannose 6-phosphate show that the position of the phosphate group of each substrate is held constant by a conserved network of hydrogen bonds. This produces two distinct, and mutually exclusive, binding orientations for the sugar rings of the 1-phospho and 6-phospho sugars. Specific binding of both orientations is accomplished by key contacts with the O3 and O4 hydroxyls of the sugar, which must occupy equatorial positions. Dual recognition of glucose and mannose phosphosugars uses a combination of specific protein contacts and nonspecific solvent contacts. The ability of PMM/PGM to accommodate these four diverse substrates in a single active site is consistent with its highly reversible phosphoryl transfer reaction and allows it to function in multiple biosynthetic pathways in P. aeruginosa.
Organizational Affiliation:
Department of Biochemistry, 117 Schweitzer Hall, University of Missouri-Columbia, Columbia, MO 65211, USA.