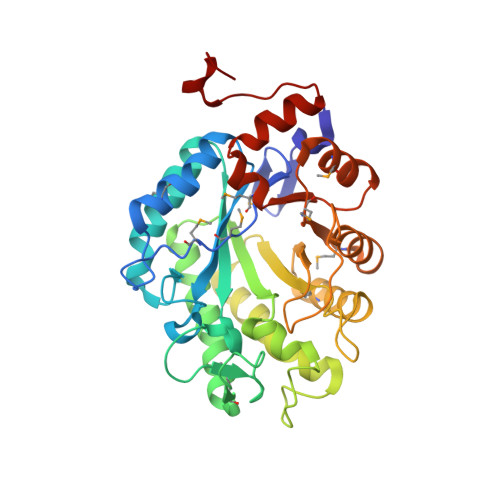
The 1.3 A Crystal Structure of the Flavoprotein YqjM Reveals a Novel Class of Old Yellow Enzymes
Kitzing, K., Fitzpatrick, T.B., Wilken, C., Sawa, J., Bourenkov, G.P., Macheroux, P., Clausen, T.(2005) J Biol Chem 280: 27904-27913
- PubMed: 15890652
- DOI: https://doi.org/10.1074/jbc.M502587200
- Primary Citation of Related Structures:
1Z41, 1Z42, 1Z44, 1Z48 - PubMed Abstract:
Here we report the crystal structure of YqjM, a homolog of Old Yellow Enzyme (OYE) that is involved in the oxidative stress response of Bacillus subtilis. In addition to the oxidized and reduced enzyme form, the structures of complexes with p-hydroxybenzaldehyde and p-nitrophenol, respectively, were solved. As for other OYE family members, YqjM folds into a (alpha/beta)8-barrel and has one molecule of flavin mononucleotide bound non-covalently at the COOH termini of the beta-sheet. Most of the interactions that control the electronic properties of the flavin mononucleotide cofactor are conserved within the OYE family. However, in contrast to all members of the OYE family characterized to date, YqjM exhibits several unique structural features. For example, the enzyme exists as a homotetramer that is assembled as a dimer of catalytically dependent dimers. Moreover, the protein displays a shared active site architecture where an arginine finger (Arg336) at the COOH terminus of one monomer extends into the active site of the adjacent monomer and is directly involved in substrate recognition. Another remarkable difference in the binding of the ligand in YqjM is represented by the contribution of the NH2-terminal Tyr28 instead of a COOH-terminal tyrosine in OYE and its homologs. The structural information led to a specific data base search from which a new class of OYE oxidoreductases was identified that exhibits a strict conservation of active site residues, which are critical for this subfamily, most notably Cys26, Tyr28, Lys109, and Arg336. Therefore, YqjM is the first representative of a new bacterial subfamily of OYE homologs.
Organizational Affiliation:
Research Institute of Molecular Pathology, Dr. Bohr-Gasse 7, A-1030 Vienna, Austria. kitzingk@gmx.net