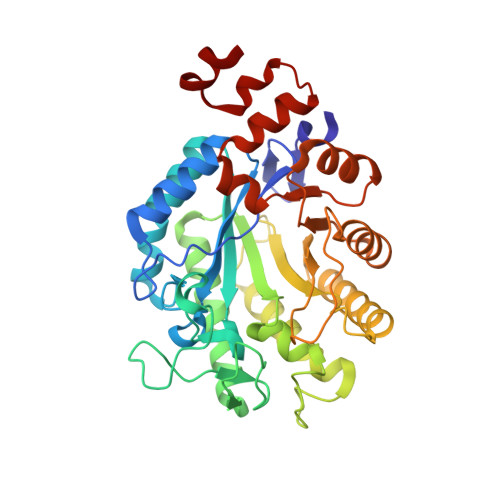
Xenobiotic reductase A in the degradation of quinoline by Pseudomonas putida 86: physiological function, structure and mechanism of 8-hydroxycoumarin reduction.
Griese, J.J., Jakob, R.P., Schwarzinger, S., Dobbek, H.(2006) J Mol Biol 361: 140-152
- PubMed: 16822524
- DOI: https://doi.org/10.1016/j.jmb.2006.06.007
- Primary Citation of Related Structures:
2H8X, 2H8Z, 2H90 - PubMed Abstract:
A continuous evolutionary pressure of the biotic and abiotic world has led to the development of a diversity of microbial pathways to degrade and biomineralize aromatic and heteroaromatic compounds. The heterogeneity of compounds metabolized by bacteria like Pseudomonas putida indicates the large variety of enzymes necessary to catalyse the required reactions. Quinoline, a N-heterocyclic aromatic compound, can be degraded by microbes along different pathways. For P. putida 86 quinoline degradation by the 8-hydroxycoumarin pathway has been described and several intermediates were identified. To select enzymes catalysing the later stages of the 8-hydroxycoumarin pathway P. putida 86 was grown with quinoline. The FMN-containing enzyme xenobiotic reductase A (XenA) was isolated and analysed for its reactivity with intermediates of the 8-hydroxycoumarin pathway. XenA catalyses the NADPH-dependent reduction of 8-hydroxycoumarin and coumarin to produce 8-hydroxy-3,4-dihydrocoumarin and 3,4-dihydrocoumarin, respectively. Crystallographic analysis of XenA alone and in complex with the two substrates revealed insights into the mechanism. XenA shows a dimeric arrangement of two (beta/alpha)(8) barrel domains each binding one FMN cofactor. High resolution crystal structures of complexes with 8-hydroxycoumarin and coumarin show different modes of binding for these molecules in the active site. While coumarin is ideally positioned for hydride transfer from N-5 of the isoalloxazine ring to C-4 of coumarin, 8-hydroxycoumarin forms a non-productive complex with oxidised XenA. Orientation of 8-hydroxycoumarin in the active site appears to be dependent on the electronic state of the flavin. We postulate that XenA catalyses the last step of the 8-hydroxycoumarin pathway before the heterocyclic ring is hydrolysed to yield 3-(2,3-dihydroxyphenyl)-propionic acid. As XenA is also found in P. putida strains unable to degrade quinoline, it appears to have more than one physiological function and is an example of how enzymes with low substrate specificity can help to explain the variety of degradation pathways possible.
Organizational Affiliation:
Laboratorium Proteinkristallographie, Universität Bayreuth, Germany.