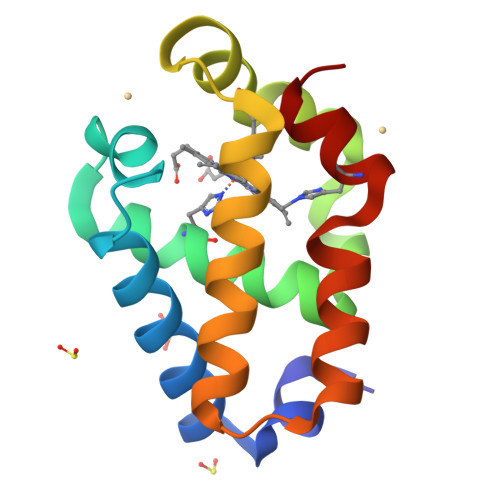
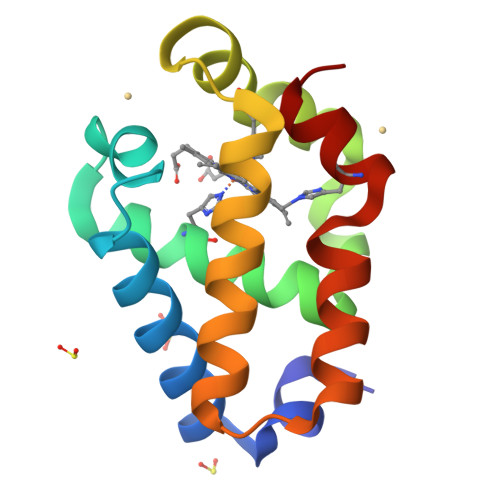
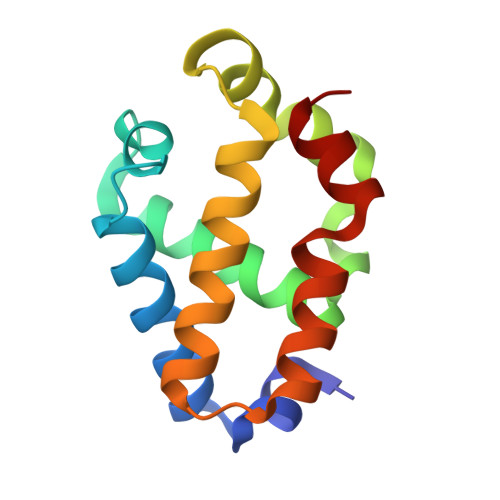
Covalent heme attachment in Synechocystis hemoglobin is required to prevent ferrous heme dissociation
Hoy, J.A., Smagghe, B.J., Halder, P., Hargrove, M.S.(2007) Protein Sci 16: 250-260
- PubMed: 17242429
- DOI: https://doi.org/10.1110/ps.062572607
- Primary Citation of Related Structures:
2HZ1, 2HZ2, 2HZ3 - PubMed Abstract:
Synechocystis hemoglobin contains an unprecedented covalent bond between a nonaxial histidine side chain (H117) and the heme 2-vinyl. This bond has been previously shown to stabilize the ferric protein against denaturation, and also to affect the kinetics of cyanide association. However, it is unclear why Synechocystis hemoglobin would require the additional degree of stabilization accompanying the His117-heme 2-vinyl bond because it also displays endogenous bis-histidyl axial heme coordination, which should greatly assist heme retention. Furthermore, the mechanism by which the His117-heme 2-vinyl bond affects ligand binding has not been reported, nor has any investigation of the role of this bond on the structure and function of the protein in the ferrous oxidation state. Here we report an investigation of the role of the Synechocystis hemoglobin His117-heme 2-vinyl bond on structure, heme coordination, exogenous ligand binding, and stability in both the ferrous and ferric oxidation states. Our results reveal that hexacoordinate Synechocystis hemoglobin lacking this bond is less stable in the ferrous oxidation state than the ferric, which is surprising in light of our understanding of pentacoordinate Hb stability, in which the ferric protein is always less stable. It is also demonstrated that removal of the His117-heme 2-vinyl bond increases the affinity constant for intramolecular histidine coordination in the ferric oxidation state, thus presenting greater competition for the ligand binding site and lowering the observed rate and affinity constants for exogenous ligands.
Organizational Affiliation:
Department of Biochemistry, Biophysics, and Molecular Biology, Iowa State University, Ames, IA 50011, USA.