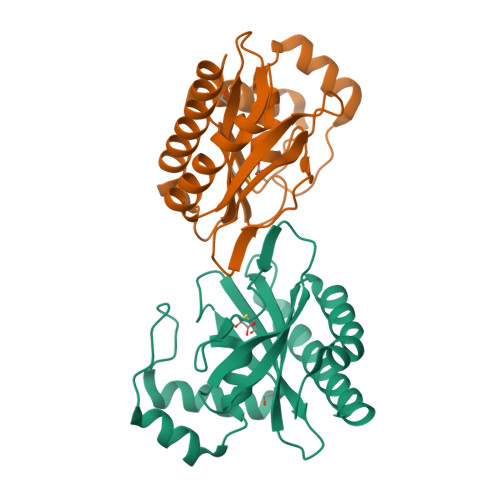
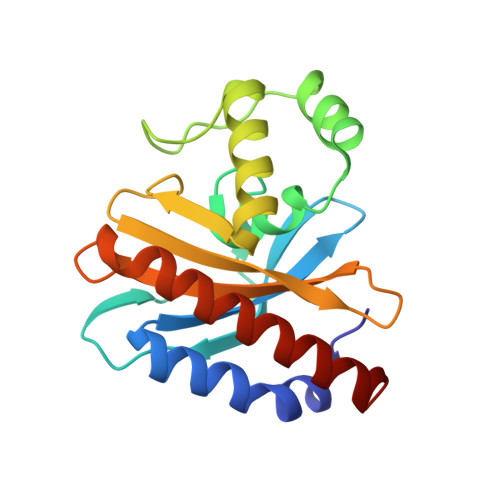
Glyoxylate and Pyruvate Are Antagonistic Effectors of the Escherichia coli IclR Transcriptional Regulator.
Lorca, G.L., Ezersky, A., Lunin, V.V., Walker, J.R., Altamentova, S., Evdokimova, E., Vedadi, M., Bochkarev, A., Savchenko, A.(2007) J Biological Chem 282: 16476-16491
- PubMed: 17426033
- DOI: https://doi.org/10.1074/jbc.M610838200
- Primary Citation of Related Structures:
2O99, 2O9A - PubMed Abstract:
The Escherichia coli isocitrate lyase regulator (IclR) regulates the expression of the glyoxylate bypass operon (aceBAK). Founding member of a large family of common fold transcriptional regulators, IclR comprises a DNA binding domain that interacts with the operator sequence and a C-terminal domain (C-IclR) that binds a hitherto unknown small molecule. We screened a chemical library of more than 150 metabolic scaffolds using a high-throughput protein stability assay to identify molecules that bind IclR and then tested the active compounds in in vitro assays of operator binding. Glyoxylate and pyruvate, identified by this method, bound the C-IclR domain with KD values of 0.9+/-0.2 and 156.2+/-7.9 microM, as defined by isothermal titration calorimetry. Both compounds altered IclR interactions with operator DNA in electrophoretic mobility shift assays but showed an antagonistic effect. Glyoxylate disrupted the formation of the IclR/operator complex in vitro by favoring the inactive dimeric state of the protein, whereas pyruvate increased the binding of IclR to the aceBAK promoter by stabilizing the active tetrameric form of the protein. Structures of the C-IclR domain alone and in complex with each effector were determined at 2.3 A, confirming the binding of both molecules in the effector recognition site previously characterized for the other representative of the family, the E. coli AllR regulator. Site-directed mutagenesis demonstrated the importance of hydrophobic patch formed by Met-146, Leu-154, Leu-220, and Leu-143 in interactions with effector molecules. In general, our strategy of combining chemical screens with functional assays and structural studies has uncovered two small molecules with antagonistic effects that regulate the IclR-dependent transcription of the aceBAK operon.
Organizational Affiliation:
Banting and Best Department of Medical Research, Toronto, Ontario M5G 1L6, Canada. glorca@ufl.edu