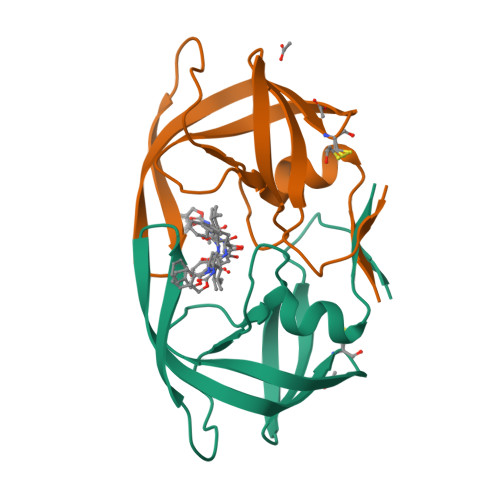
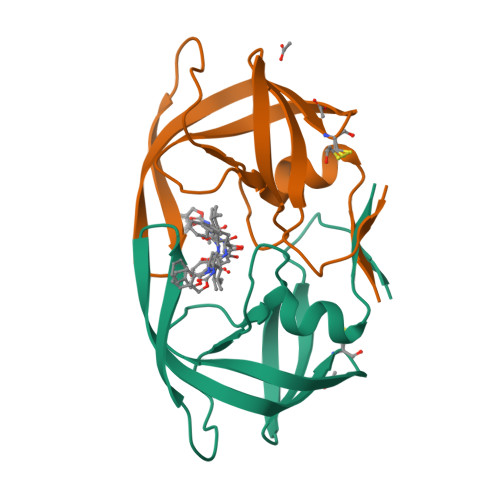
Structural Characterization of B and non-B Subtypes of HIV-Protease: Insights into the Natural Susceptibility to Drug Resistance Development.
Sanches, M., Krauchenco, S., Martins, N.H., Gustchina, A., Wlodawer, A., Polikarpov, I.(2007) J Mol Biology 369: 1029-1040
- PubMed: 17467738
- DOI: https://doi.org/10.1016/j.jmb.2007.03.049
- Primary Citation of Related Structures:
2P3A, 2P3B, 2P3C, 2P3D - PubMed Abstract:
Although a majority of HIV-1 infections in Brazil are caused by the subtype B virus (also prevalent in the United States and Western Europe), viral subtypes F and C are also found very frequently. Genomic differences between the subtypes give rise to sequence variations in the encoded proteins, including the HIV-1 protease. The current anti-HIV drugs have been developed primarily against subtype B and the effects arising from the combination of drug-resistance mutations with the naturally existing polymorphisms in non-B HIV-1 subtypes are only beginning to be elucidated. To gain more insights into the structure and function of different variants of HIV proteases, we have determined a 2.1 A structure of the native subtype F HIV-1 protease (PR) in complex with the protease inhibitor TL-3. We have also solved crystal structures of two multi-drug resistant mutant HIV PRs in complex with TL-3, from subtype B (Bmut) carrying the primary mutations V82A and L90M, and from subtype F (Fmut) carrying the primary mutation V82A plus the secondary mutation M36I, at 1.75 A and 2.8 A resolution, respectively. The proteases Bmut, Fwt and Fmut exhibit sevenfold, threefold, and 54-fold resistance to TL-3, respectively. In addition, the structure of subtype B wild type HIV-PR in complex with TL-3 has been redetermined in space group P6(1), consistent with the other three structures. Our results show that the primary mutation V82A causes the known effect of collapsing the S1/S1' pockets that ultimately lead to the reduced inhibitory effect of TL-3. Our results further indicate that two naturally occurring polymorphic substitutions in subtype F and other non-B HIV proteases, M36I and L89M, may lead to early development of drug resistance in patients infected with non-B HIV subtypes.
Organizational Affiliation:
Grupo de Cristalografia, Instituto de Física de São Carlos, Universidade de São Paulo, Av. Trabalhador Saocarlense, 400, CEP 13560-970, São Carlos, SP, Brazil.