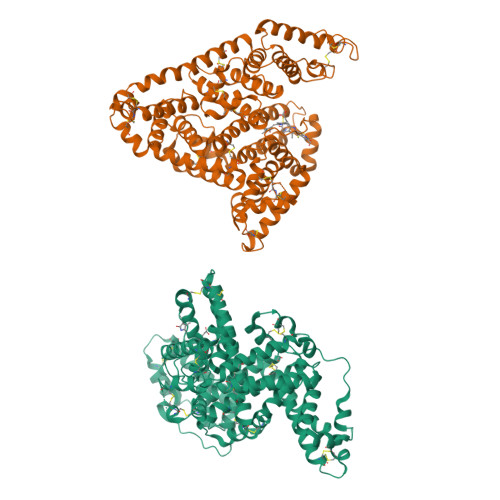
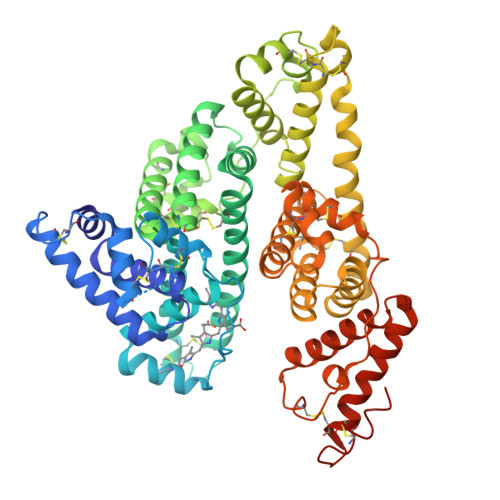
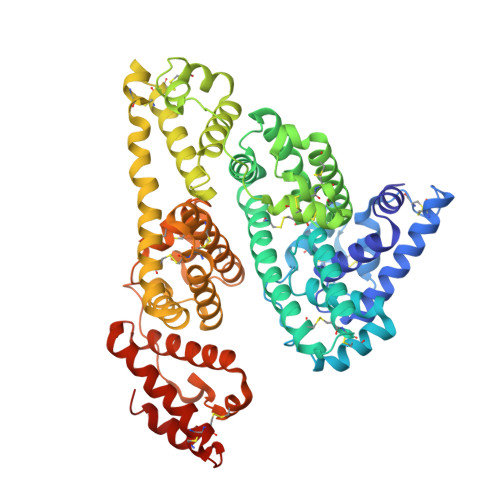
Crystallographic Analysis of Human Serum Albumin Complexed with 4Z,15E-Bilirubin-Ixalpha.
Zunszain, P.A., Ghuman, J., Mcdonagh, A.F., Curry, S.(2008) J Mol Biology 381: 394
- PubMed: 18602119
- DOI: https://doi.org/10.1016/j.jmb.2008.06.016
- Primary Citation of Related Structures:
2VUE, 2VUF - PubMed Abstract:
Bilirubin, an insoluble yellow-orange pigment derived from heme catabolism, accumulates to toxic levels in individuals with impaired or immature liver function. The resulting jaundice may be managed with phototherapy to isomerize the biosynthetic 4Z,15Z-bilirubin-IXalpha to more soluble and excretable isomers, such as 4Z,15E-bilirubin. Bilirubin and its configurational isomers are transported to the liver by human serum albumin (HSA) but their precise binding location(s) on the protein have yet to be determined. To investigate the molecular details of their interaction, we co-crystallised bilirubin with HSA. Strikingly, the crystal structure--determined to 2.42 A resolution--revealed the 4Z,15E-bilirubin-IXalpha isomer bound to an L-shaped pocket in sub-domain IB. We also determined the co-crystal structure of HSA complexed with fusidic acid, an antibiotic that competitively displaces bilirubin from the protein, and showed that it binds to the same pocket. These results provide the first crystal structure of a natural bilirubin pigment bound to serum albumin, challenge some of the present conceptions about HSA-bilirubin interactions, and provide a sound structural framework for finally resolving the long-standing question of where 4Z,15Z-bilirubin-IXalpha binds to the protein.
- Biophysics Section, Blackett Laboratory, Imperial College, South Kensington Campus, Exhibition Road, London SW7 2AZ, UK.
Organizational Affiliation: