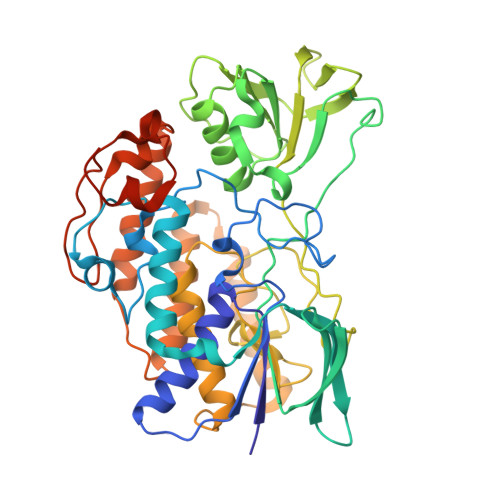
Structural and Functional Analysis of Bacterial Flavin-Containing Monooxygenase Reveals its Ping-Pong-Type Reaction Mechanism.
Cho, H.J., Cho, H.Y., Kim, K.J., Kim, M.H., Kim, S.W., Kang, B.S.(2011) J Struct Biol 175: 39
- PubMed: 21527346
- DOI: https://doi.org/10.1016/j.jsb.2011.04.007
- Primary Citation of Related Structures:
2XVE, 2XVF, 2XVH, 2XVI, 2XVJ - PubMed Abstract:
A bacterial flavin-containing monooxygenase (bFMO) catalyses the oxygenation of indole to produce indigoid compounds. In the reductive half of the indole oxygenation reaction, NADPH acts as a reducing agent, and NADP(+) remains at the active site, protecting bFMO from reoxidation. Here, the crystal structures of bFMO and bFMO in complex with NADP(+), and a mutant bFMO(Y207S), which lacks indole oxygenation activity, with and without indole are reported. The crystal structures revealed overlapping binding sites for NADP(+) and indole, suggestive of a double-displacement reaction mechanism for bFMO. In biochemical assays, indole inhibited NADPH oxidase activity, and NADPH in turn inhibited the binding of indole and decreased indoxyl production. Comparison of the structures of bFMO with and without bound NADP(+) revealed that NADPH induces conformational changes in two active site motifs. One of the motifs contained Arg-229, which participates in interactions with the phosphate group of NADPH and appears be a determinant of the preferential binding of bFMO to NADPH rather than NADH. The second motif contained Tyr-207. The mutant bFMO(Y207S) exhibited very little indoxyl producing activity; however, the NADPH oxidase activity of the mutant was higher than the wild-type enzyme. It suggests a role for Y207, in the protection of hydroperoxyFAD. We describe an indole oxygenation reaction mechanism for bFMO that involves a ping-pong-like interaction of NADPH and indole.
Organizational Affiliation:
School of Life Science and Biotechnology, Kyungpook National University, Buk-gu, Daegu 702-701, Republic of Korea.