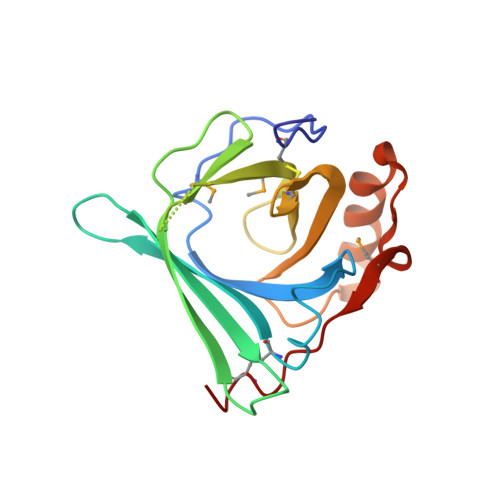
Structures of two major allergens, Bla g 4 and Per a 4, from cockroaches and their IgE binding epitopes.
Tan, Y.W., Chan, S.L., Ong, T.C., Yit, L.Y., Tiong, Y.S., Chew, F.T., Sivaraman, J., Mok, Y.K.(2008) J Biol Chem 284: 3148-3157
- PubMed: 19056737
- DOI: https://doi.org/10.1074/jbc.M807209200
- Primary Citation of Related Structures:
3EBK, 3EBW - PubMed Abstract:
Inhalant allergens from cockroaches are an important cause of asthma to millions of individuals worldwide. Here we report for the first time the structures of two major cockroach allergens, Bla g 4 and Per a 4, that adopt a typical lipocalin fold but with distinct structural features as compared with other known lipocalin allergens. Both Bla g 4 and Per a 4 contain two long-range disulfide bonds linking the N and C termini to a beta-barrel. The C-terminal helix of Bla g 4 is bent and greatly extended toward the N terminus. Bla g 4 is found to be a monomer, whereas Per a 4 exists as a dimer in solution with a novel dimeric interface involving residues from loops at the top and bottom of the beta-barrel. Putative ligand binding sites of both allergens are determined by docking of the juvenile hormone III inside the beta-barrel and found to interact with the ligand using non-conserved residues. Bla g 4 and Per a 4 are found to be cross-reactive in sera IgE binding, at least in the Singaporean Chinese population tested. A major IgE binding epitope unique to Per a 4 is found on the loops at the bottom of the beta-barrel that may aid the development of hypoallergens for immunotherapy.
Organizational Affiliation:
Department of Biological Sciences, National University of Singapore, Singapore 17543.