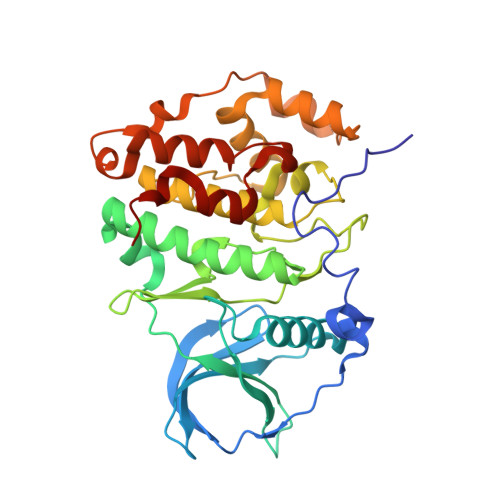
Structural basis of CX-4945 binding to human protein kinase CK2.
Ferguson, A.D., Sheth, P.R., Basso, A.D., Paliwal, S., Gray, K., Fischmann, T.O., Le, H.V.(2011) FEBS Lett 585: 104-110
- PubMed: 21093442
- DOI: https://doi.org/10.1016/j.febslet.2010.11.019
- Primary Citation of Related Structures:
3NGA, 3NSZ - PubMed Abstract:
Protein kinase CK2 (CK2), a constitutively active serine/threonine kinase, is involved in a variety of roles essential to the maintenance of cellular homeostasis. Elevated levels of CK2 expression results in the dysregulation of key signaling pathways that regulate transcription, and has been implicated in cancer. The adenosine-5'-triphosphate-competitive inhibitor CX-4945 has been reported to show broad spectrum anti-proliferative activity in multiple cancer cell lines. Although the enzymatic IC(50) of CX-4945 has been reported, the thermodynamics and structural basis of binding to CK2α remained elusive. Presented here are the crystal structures of human CK2α in complex with CX-4945 and adenylyl phosphoramidate at 2.7 and 1.3 Å, respectively. Biophysical analysis of CX-4945 binding is also described. This data provides the structural rationale for the design of more potent inhibitors against this emerging cancer target.
Organizational Affiliation:
Drug Design Department, Merck Research Laboratory, Kenilworth, NJ 07033, USA. andrew.ferguson@astrazeneca.com