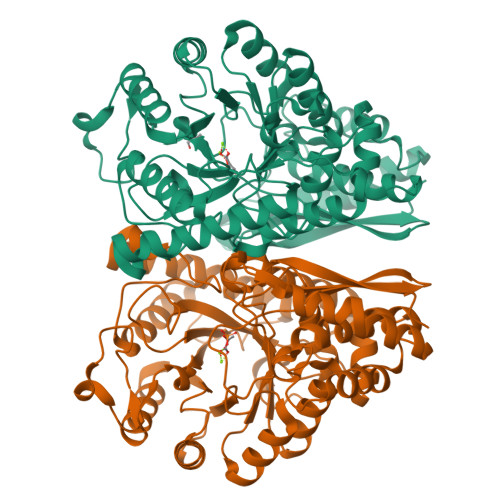
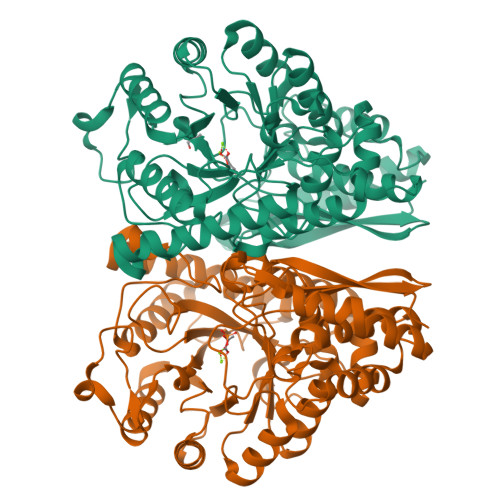
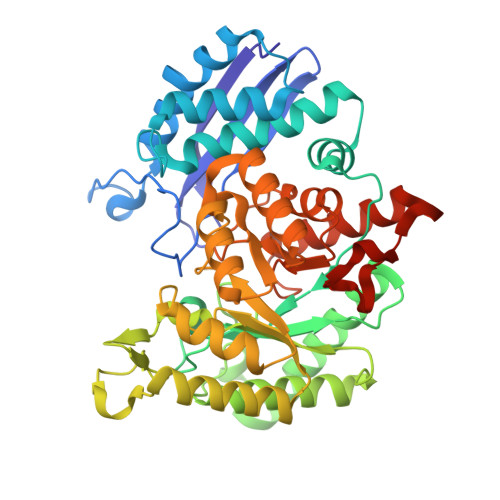
Structures of asymmetric complexes of human neuron specific enolase with resolved substrate and product and an analogous complex with two inhibitors indicate subunit interaction and inhibitor cooperativity.
Qin, J., Chai, G., Brewer, J.M., Lovelace, L.L., Lebioda, L.(2012) J Inorg Biochem 111: 187-194
- PubMed: 22437160
- DOI: https://doi.org/10.1016/j.jinorgbio.2012.02.011
- Primary Citation of Related Structures:
3UCC, 3UCD, 3UJE, 3UJF, 3UJR, 3UJS - PubMed Abstract:
In the presence of magnesium, enolase catalyzes the dehydration of 2-phospho-d-glycerate (PGA) to phosphoenolpyruvate (PEP) in glycolysis and the reverse reaction in gluconeogensis at comparable rates. The structure of human neuron specific enolase (hNSE) crystals soaked in PGA showed that the enzyme is active in the crystals and produced PEP; conversely soaking in PEP produced PGA. Moreover, the hNSE dimer contains PGA bound in one subunit and PEP or a mixture of PEP and PGA in the other. Crystals soaked in a mixture of competitive inhibitors tartronate semialdehyde phosphate (TSP) and lactic acid phosphate (LAP) showed asymmetry with TSP binding in the same site as PGA and LAP in the PEP site. Kinetic studies showed that the inhibition of NSE by mixtures of TSP and LAP is stronger than predicted for independently acting inhibitors. This indicates that in some cases inhibition of homodimeric enzymes by mixtures of inhibitors ("heteroinhibition") may offer advantages over single inhibitors.
Organizational Affiliation:
Department of Chemistry & Biochemistry, University of South Carolina, Columbia, SC 29208, United States.