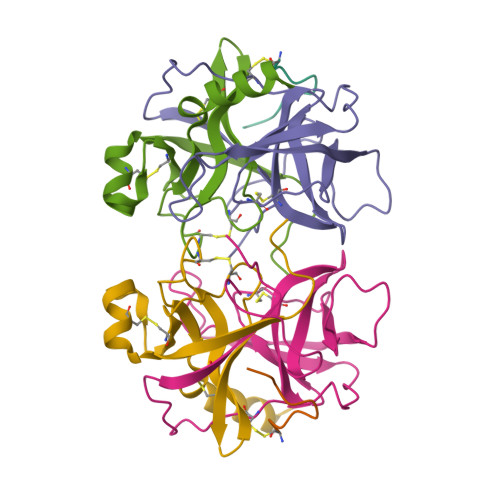
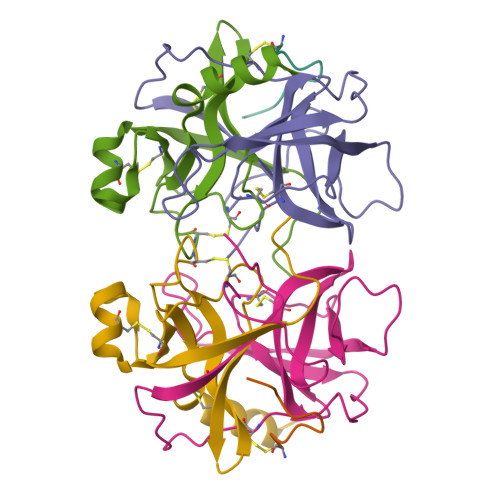
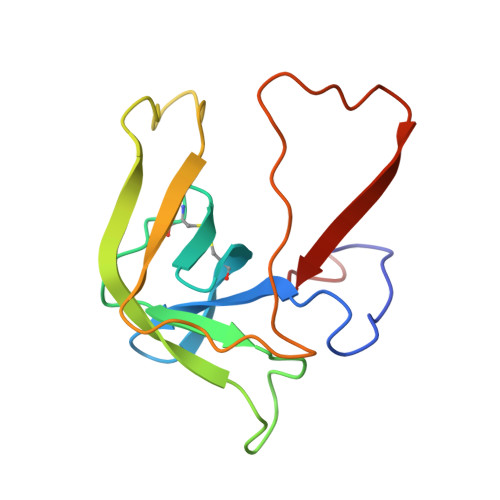
Structure of alpha-chymotrypsin refined at 1.68 A resolution.
Tsukada, H., Blow, D.M.(1985) J Mol Biology 184: 703-711
- PubMed: 4046030
- DOI: https://doi.org/10.1016/0022-2836(85)90314-6
- Primary Citation of Related Structures:
4CHA - PubMed Abstract:
Diffraction data for alpha-chymotrypsin crystals at -10 degrees C were measured at 1.68 A resolution and refined by restrained structure-factor least-squares refinement. The two independent chymotrypsin molecules in the crystallographic asymmetric unit were refined independently. The overall structure of alpha-chymotrypsin is little changed from published co-ordinates. The root-mean-square shift of C alpha co-ordinates is 0.42 A, co-ordinates for the two molecules showing a root-mean-square difference of 0.19 A. Certain regions with high disorder (residues 9 to 14, 73 to 79) remain difficult to interpret and several side-chains are disordered. Some water molecule positions have been changed. The absence of the tosyl group has made a significant difference to the refined structure at the active site. This now agrees closely with other enzymes of the trypsin family that have been refined at high resolution. There is a strong hydrogen bond between N epsilon 2 (His57) and O gamma (Ser195) in the free enzyme, in line with the published description of the charge relay system.