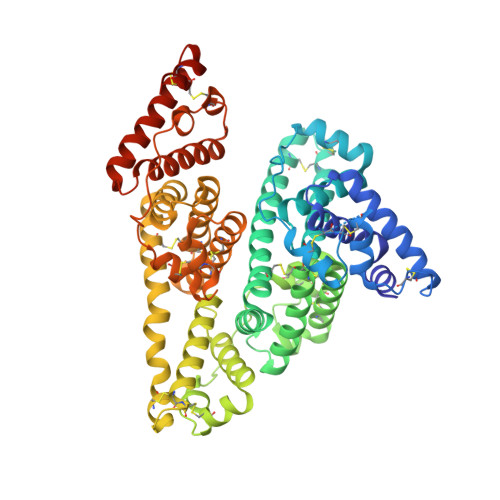
Structural evidence of perfluorooctane sulfonate transport by human serum albumin
Luo, Z.P., Shi, X.L., Hu, Q., Zhao, B., Huang, M.D.(2012) Chem Res Toxicol 25: 990-992
- PubMed: 22482699
- DOI: https://doi.org/10.1021/tx300112p
- Primary Citation of Related Structures:
4E99 - PubMed Abstract:
Perfluorooctane sulfonate (PFOS) is a man-made fluorosurfactant and globally persistent organic pollutant. PFOS is mainly distributed in blood with a long half-life for elimination. PFOS was found mainly bound to human serum albumin (HSA) in plasma, the most abundant protein in human blood plasma, which transports a variety of endogenous and exogenous ligands. However, the structural basis of such binding remains unclear. Here, we report the crystal structure of the HSA-PFOS complex and show that PFOS binds to HSA at a molar ratio of 2:1. In addition, PFOS binding renders the HSA structure more compact. Our results provide a structural mechanism to understand the retention of surfactants in human serum.
Organizational Affiliation:
State Key laboratory of Structural Chemistry, Fujian Institute of Research on the Structure of Matter, Chinese Academy of Sciences, Fuzhou, China.