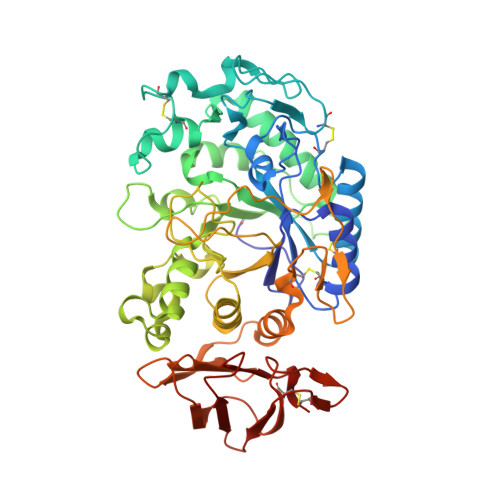
Order and disorder: differential structural impacts of myricetin and ethyl caffeate on human amylase, an antidiabetic target.
Williams, L.K., Li, C., Withers, S.G., Brayer, G.D.(2012) J Med Chem 55: 10177-10186
- PubMed: 23050660
- DOI: https://doi.org/10.1021/jm301273u
- Primary Citation of Related Structures:
4GQQ, 4GQR - PubMed Abstract:
The increasing prevalence of diabetes has accelerated the search for new drugs derived from natural sources. To define the functional features of two such families of compounds, the flavonols and the ethyl caffeates, we have determined the high-resolution structures of representative inhibitors in complex with human pancreatic α-amylase. Myricetin binds at the active site and interacts directly with the catalytic residues despite its bulky planar nature. Notably, it reduces the normal conformational flexibility of the adjacent substrate binding cleft. In contrast, bound ethyl caffeate acts by disordering precisely those polypeptide chain segments that make up the active site binding cleft. It also operates from binding sites far removed from the active site, a property not observed in any other class of human α-amylase inhibitor studied to date. Given the current inadequacy of drugs directed at diabetes, the use of optimized flavonols and ethyl caffeates may present an alternative therapeutic route.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, University of British Columbia , Vancouver, British Columbia V6T 1Z3, Canada.