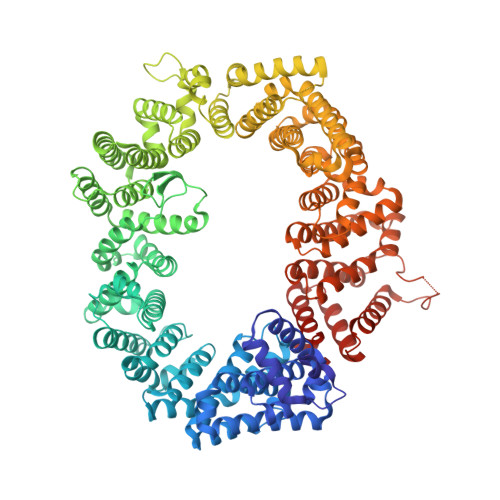
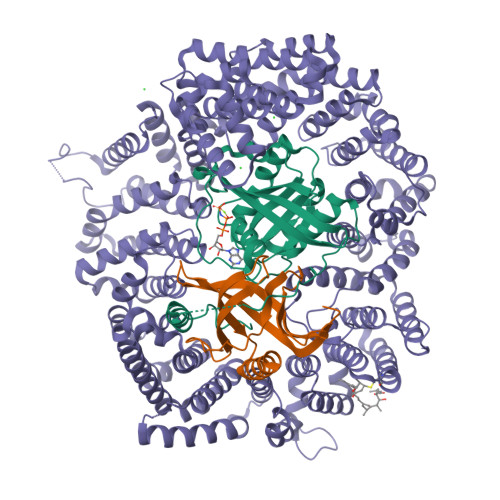
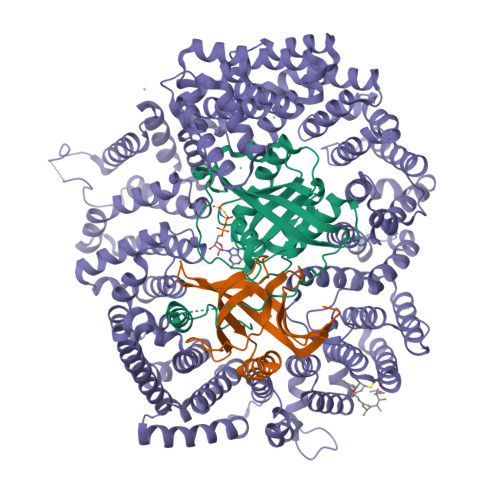
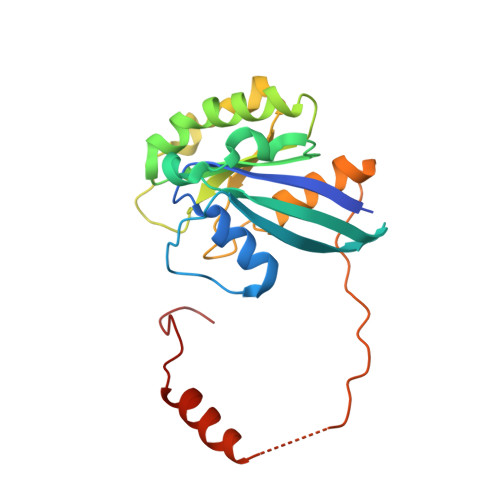
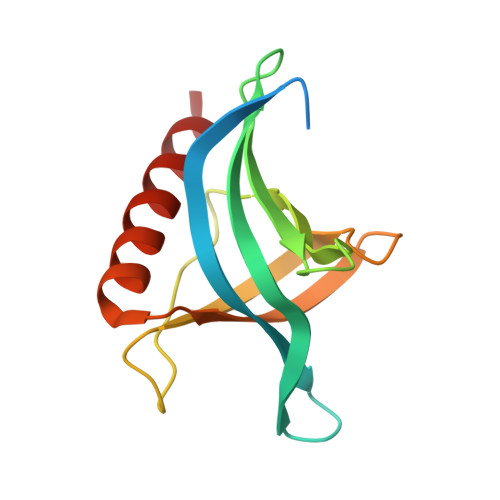
Nuclear export inhibition through covalent conjugation and hydrolysis of Leptomycin B by CRM1.
Sun, Q., Carrasco, Y.P., Hu, Y., Guo, X., Mirzaei, H., Macmillan, J., Chook, Y.M.(2013) Proc Natl Acad Sci U S A 110: 1303-1308
- PubMed: 23297231
- DOI: https://doi.org/10.1073/pnas.1217203110
- Primary Citation of Related Structures:
4HAT, 4HAU, 4HAV, 4HAW, 4HAX, 4HAY, 4HAZ, 4HB0, 4HB2, 4HB3, 4HB4 - PubMed Abstract:
The polyketide natural product Leptomycin B inhibits nuclear export mediated by the karyopherin protein chromosomal region maintenance 1 (CRM1). Here, we present 1.8- to 2.0-Å-resolution crystal structures of CRM1 bound to Leptomycin B and related inhibitors Anguinomycin A and Ratjadone A. Structural and complementary chemical analyses reveal an unexpected mechanism of inhibition involving covalent conjugation and CRM1-mediated hydrolysis of the natural products' lactone rings. Furthermore, mutagenesis reveals the mechanism of hydrolysis by CRM1. The nuclear export signal (NES)-binding groove of CRM1 is able to drive a chemical reaction in addition to binding protein cargoes for transport through the nuclear pore complex.
Organizational Affiliation:
Department of Pharmacology, University of Texas Southwestern Medical Center, Dallas, TX 75390-9041, USA.