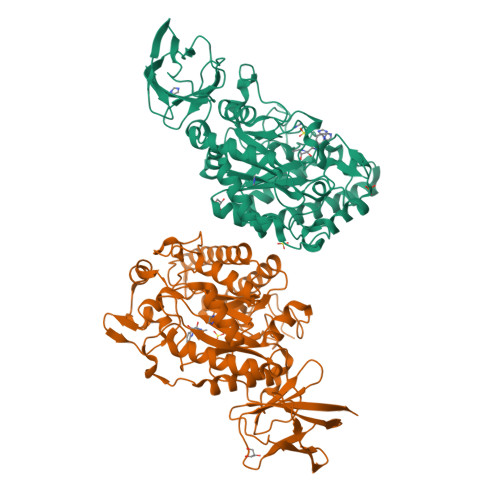
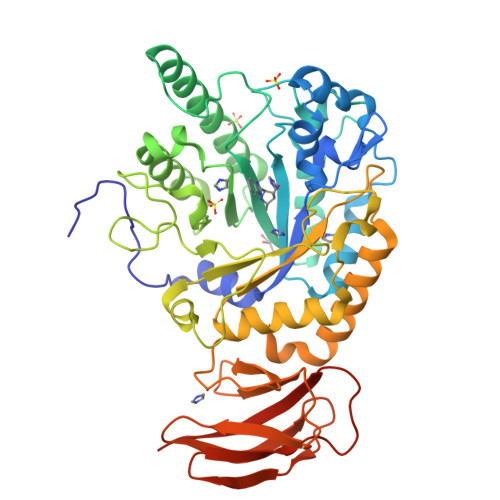
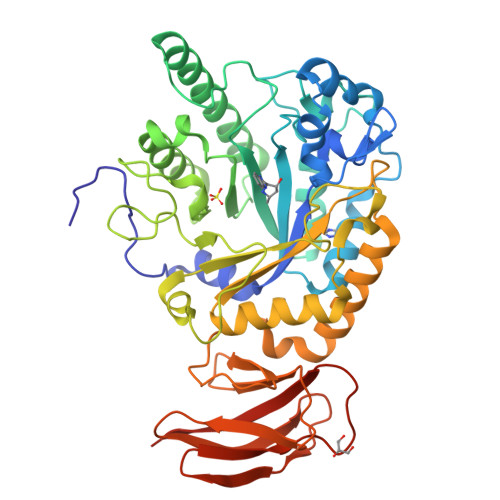
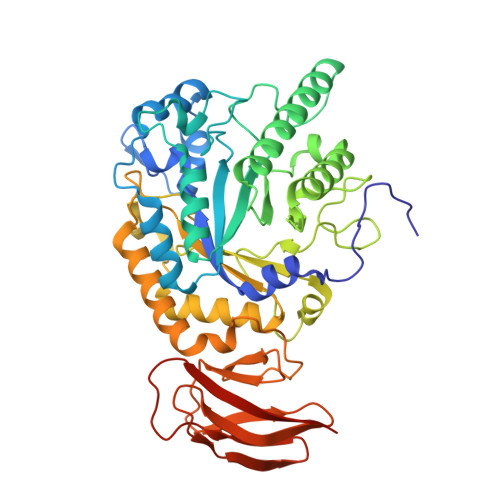
Three dimensional structure of a bacterial alpha-l-fucosidase with a 5-membered iminocyclitol inhibitor.
Wright, D.W., Moreno-Vargas, A.J., Carmona, A.T., Robina, I., Davies, G.J.(2013) Bioorg Med Chem 21: 4751-4754
- PubMed: 23830696
- DOI: https://doi.org/10.1016/j.bmc.2013.05.056
- Primary Citation of Related Structures:
4J27, 4J28 - PubMed Abstract:
Fucosidases, enzymes that cleave fucose from the non-reducing end of a glycan, represent promising medicinal targets reflecting their roles in cancer metastasis, inflammation, host-parasite interactions and the lysosomal storage disorder fucosidosis. The X-ray crystal structures of Bacteroides thetaiotaomicron GH29 α-l-fucosidase (BtFuc2970) in a new crystal form (at a resolution of 1.59Å) and liganded with a 5-membered iminocyclitol inhibitor (1.73Å) are reported herein. The 5-membered iminocyclitol binds in a (3)E conformation, mimicking the proposed (3)H4 half chair transition-state of the enzyme catalysed reaction, and its Ki for BtFuc2970 was determined as 2μM. Structural analysis of fucosidase inhibition through 5-membered iminocyclitols will aid in the rational design of more potent fucosidase inhibitors for treatment of a range of medical conditions.
Organizational Affiliation:
Structural Biology Laboratory, Department of Chemistry, University of York, York YO10 5DD, United Kingdom.