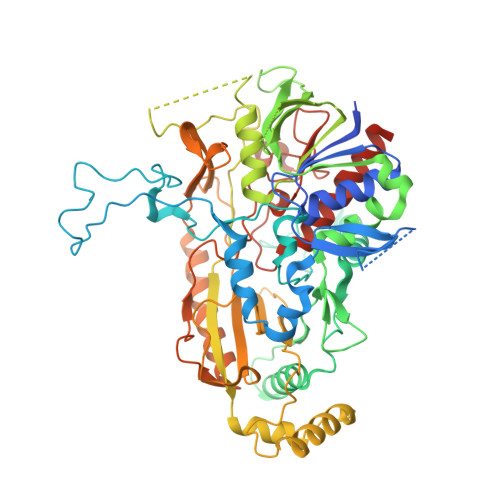
Crystal structures of Phanerochaete chrysosporium pyranose 2-oxidase suggest that the N-terminus acts as a propeptide that assists in homotetramer assembly.
Hassan, N., Tan, T.C., Spadiut, O., Pisanelli, I., Fusco, L., Haltrich, D., Peterbauer, C.K., Divne, C.(2013) FEBS Open Bio 3: 496-504
- PubMed: 24282677
- DOI: https://doi.org/10.1016/j.fob.2013.10.010
- Primary Citation of Related Structures:
4MIF, 4MIG, 4MIH - PubMed Abstract:
The flavin-dependent homotetrameric enzyme pyranose 2-oxidase (P2O) is found mostly, but not exclusively, in lignocellulose-degrading fungi where it catalyzes the oxidation of β-d-glucose to the corresponding 2-keto sugar concomitantly with hydrogen peroxide formation during lignin solubilization. Here, we present crystal structures of P2O from the efficient lignocellulolytic basidiomycete Phanerochaete chrysosporium. Structures were determined of wild-type PcP2O from the natural fungal source, and two variants of recombinant full-length PcP2O, both in complex with the slow substrate 3-deoxy-3-fluoro-β-d-glucose. The active sites in PcP2O and P2O from Trametes multicolor (TmP2O) are highly conserved with identical substrate binding. Our structural analysis suggests that the 17 °C higher melting temperature of PcP2O compared to TmP2O is due to an increased number of intersubunit salt bridges. The structure of recombinant PcP2O expressed with its natural N-terminal sequence, including a proposed propeptide segment, reveals that the first five residues of the propeptide intercalate at the interface between A and B subunits to form stabilizing, mainly hydrophobic, interactions. In the structure of mature PcP2O purified from the natural source, the propeptide segment in subunit A has been replaced by a nearby loop in the B subunit. We propose that the propeptide in subunit A stabilizes the A/B interface of essential dimers in the homotetramer and that, upon maturation, it is replaced by the loop in the B subunit to form the mature subunit interface. This would imply that the propeptide segment of PcP2O acts as an intramolecular chaperone for oligomerization at the A/B interface of the essential dimer.
Organizational Affiliation:
KTH Royal Institute of Technology, School of Biotechnology, Albanova University Center, Roslagstullsbacken 21, S-10691 Stockholm Sweden.