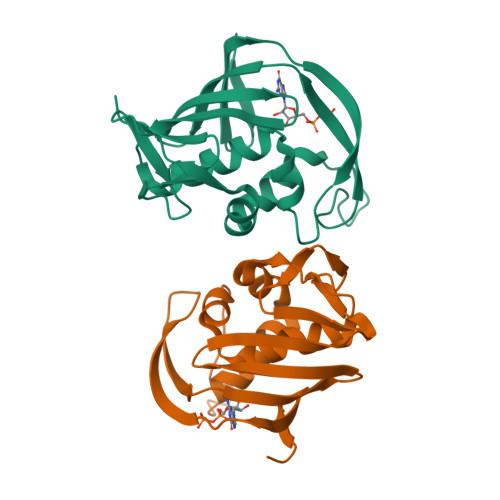
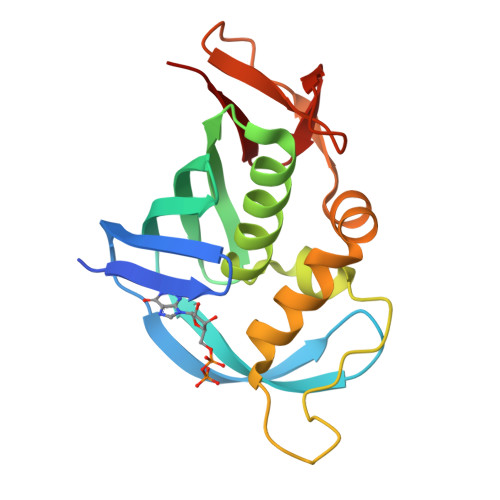
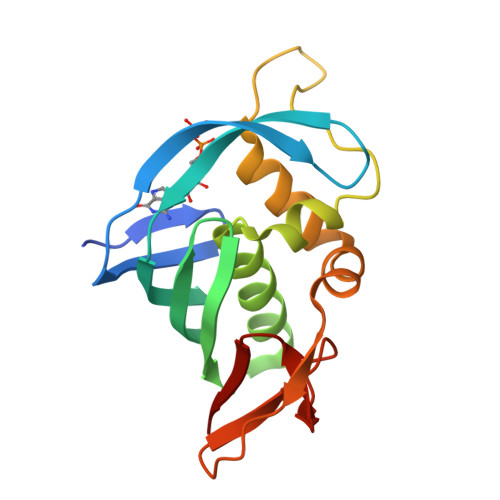
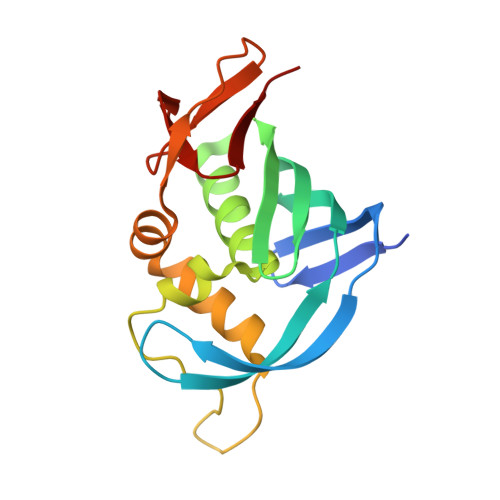
The crystal structure of the PB2 cap-binding domain of influenza B virus reveals a novel cap recognition mechanism.
Liu, Y., Yang, Y., Fan, J., He, R., Luo, M., Zheng, X.(2015) J Biological Chem 290: 9141-9149
- PubMed: 25691568
- DOI: https://doi.org/10.1074/jbc.M115.636464
- Primary Citation of Related Structures:
4OR4, 4OR6, 4Q46 - PubMed Abstract:
The influenza RNA-dependent RNA polymerase is a core enzyme required for both transcription and replication of the virus RNA genome, making it a potential drug target for the influenza virus. To detect the feature of cap-dependent transcription of influenza B virus (FluB) polymerase, we determined the crystal structures of the wild-type FluB polymerase PB2 subunit cap-binding domain (PB2cap) with bound GDP and the mutant FluB Q325F PB2cap with bound m(7)GDP or GDP. These structures revealed that, distinct from influenza A virus (FluA) PB2cap, the guanine and ribose moieties of substrates invert in FluB PB2caps. Moreover, we characterized the substrate specificity and affinity of the PB2caps using isothermal titration calorimetry. FluB PB2cap has a weaker affinity for m(7)GDP than FluA PB2cap. Unlike FluA PB2cap that has a preference for m(7)GDP in comparison with GDP, FluB PB2cap shows an analogous affinity for both substrates. Replacement of FluB PB2 Glu(325) by Phe, the corresponding residue of FluA PB2, increased the binding affinity of FluB PB2cap for m(7)GDP to a level approximate to that of FluA PB2cap and caused a significant higher affinity to GDP. This study indicated that FluB PB2cap has a unique cap recognition mechanism compared with FluA PB2cap, providing molecular insight into inhibitor design targeting FluB PB2cap.
Organizational Affiliation:
From the State Key Lab of Protein and Plant Gene Research and Department of Biochemistry and Molecular Biology, School of Life Sciences, Peking University, Beijing 100871, China and.