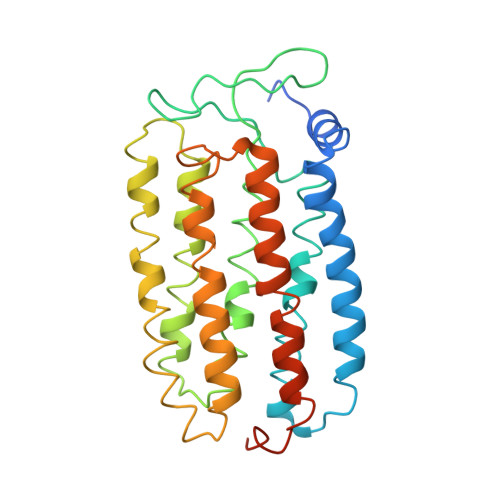
Crystal Structures of the L1, L2, N, and O States of pharaonis Halorhodopsin
Kouyama, T., Kawaguchi, H., Nakanishi, T., Kubo, H., Murakami, M.(2015) Biophys J 108: 2680-2690
- PubMed: 26039169
- DOI: https://doi.org/10.1016/j.bpj.2015.04.027
- Primary Citation of Related Structures:
4QRY - PubMed Abstract:
Halorhodopsin from Natronomonas pharaonis (pHR) functions as a light-driven halide ion pump. In the presence of halide ions, the photochemical reaction of pHR is described by the scheme: K→ L1 → L2 → N → O → pHR' → pHR. Here, we report light-induced structural changes of the pHR-bromide complex observed in the C2 crystal. In the L1-to-L2 transition, the bromide ion that initially exists in the extracellular vicinity of retinal moves across the retinal Schiff base. Upon the formation of the N state with a bromide ion bound to the cytoplasmic vicinity of the retinal Schiff base, the cytoplasmic half of helix F moves outward to create a water channel in the cytoplasmic interhelical space, whereas the extracellular half of helix C moves inward. During the transition from N to an N-like reaction state with retinal assuming the 13-cis/15-syn configuration, the translocated bromide ion is released into the cytoplasmic medium. Subsequently, helix F relaxes into its original conformation, generating the O state. Anion uptake from the extracellular side occurs when helix C relaxes into its original conformation. These structural data provide insight into the structural basis of unidirectional anion transport.
Organizational Affiliation:
Department of Physics, Graduate School of Science, Nagoya University, Nagoya, Japan; RIKEN Harima Branch, 1-1-1, Kouto, Sayo, Hyogo, Japan. Electronic address: kouyama@bio.phys.nagoya-u.ac.jp.