Optimization of 5-(2,6-dichlorophenyl)-3-hydroxy-2-mercaptocyclohex-2-enones as potent inhibitors of human lactate dehydrogenase.
Labadie, S., Dragovich, P.S., Chen, J., Fauber, B.P., Boggs, J., Corson, L.B., Ding, C.Z., Eigenbrot, C., Ge, H., Ho, Q., Lai, K.W., Ma, S., Malek, S., Peterson, D., Purkey, H.E., Robarge, K., Salphati, L., Sideris, S., Ultsch, M., VanderPorten, E., Wei, B., Xu, Q., Yen, I., Yue, Q., Zhang, H., Zhang, X., Zhou, A.(2014) Bioorg Med Chem Lett 25: 75-82
- PubMed: 25466195
- DOI: https://doi.org/10.1016/j.bmcl.2014.11.008
- Primary Citation of Related Structures:
4R68, 4R69 - PubMed Abstract:
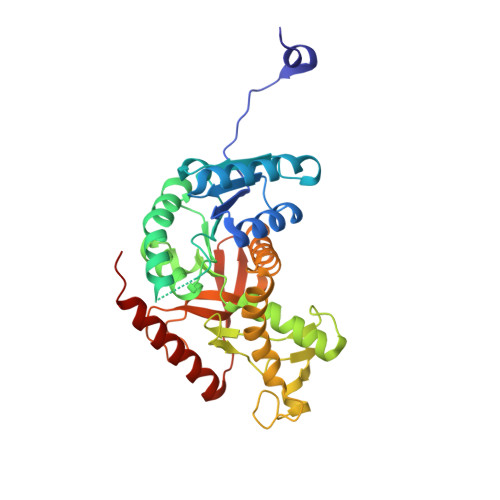
Optimization of 5-(2,6-dichlorophenyl)-3-hydroxy-2-mercaptocyclohex-2-enone using structure-based design strategies resulted in inhibitors with considerable improvement in biochemical potency against human lactate dehydrogenase A (LDHA). These potent inhibitors were typically selective for LDHA over LDHB isoform (4–10 fold) and other structurally related malate dehydrogenases, MDH1 and MDH2 (>500 fold). An X-ray crystal structure of enzymatically most potent molecule bound to LDHA revealed two additional interactions associated with enhanced biochemical potency.
Organizational Affiliation:
Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, USA.