Characterization and crystal structure determination of beta-1,2-mannobiose phosphorylase from Listeria innocua
Tsuda, T., Nihira, T., Chiku, K., Suzuki, E., Arakawa, T., Nishimoto, M., Kitaoka, M., Nakai, H., Fushinobu, S.(2015) FEBS Lett 589: 3816-3821
- PubMed: 26632508
- DOI: https://doi.org/10.1016/j.febslet.2015.11.034
- Primary Citation of Related Structures:
5B0P, 5B0Q, 5B0R, 5B0S - PubMed Abstract:
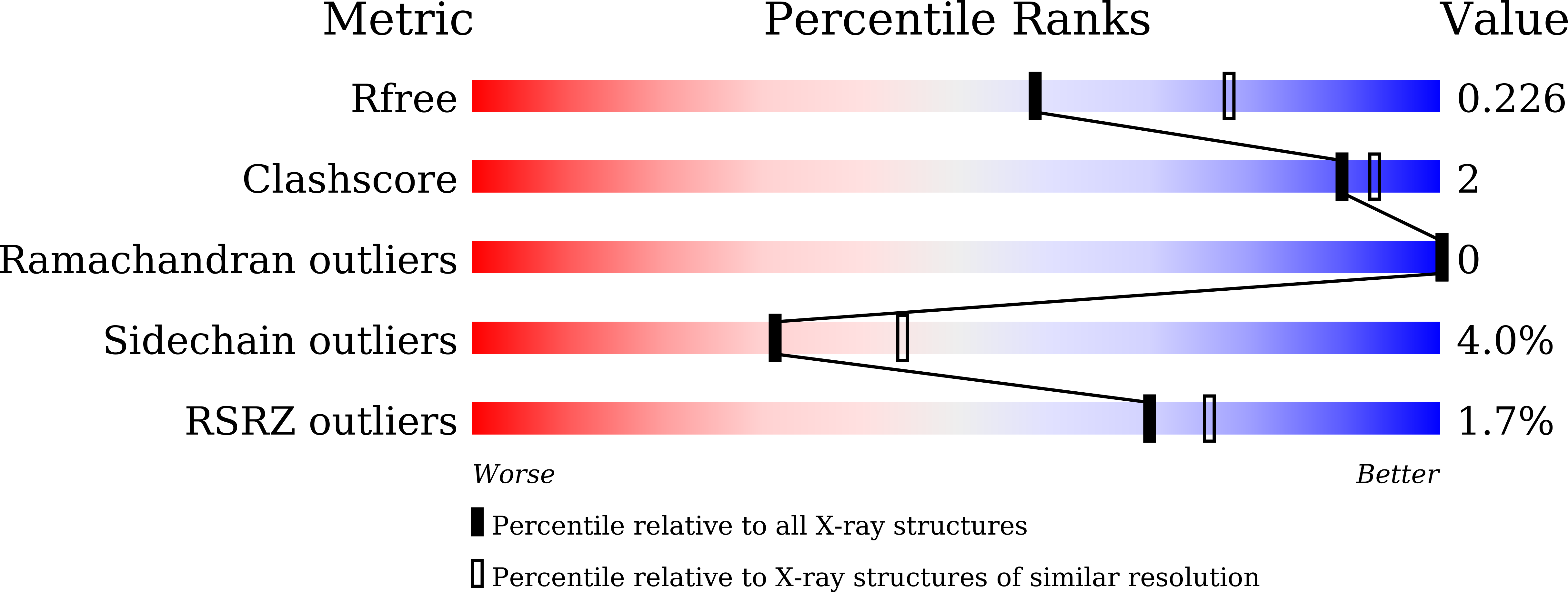
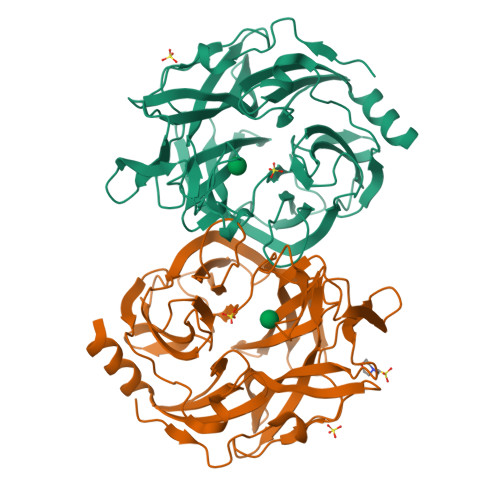
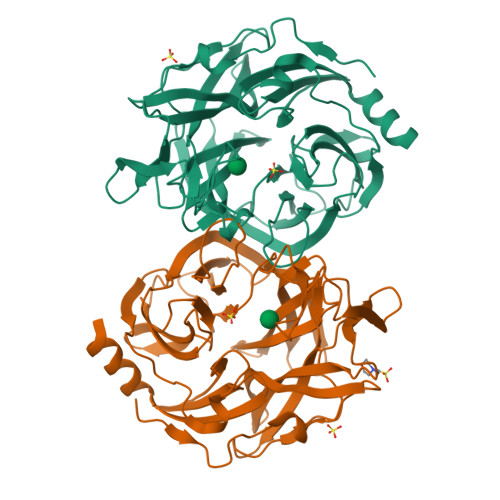
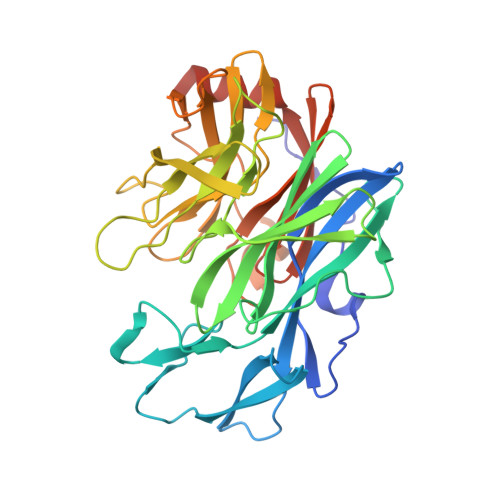
Glycoside hydrolase family 130 consists of phosphorylases and hydrolases for β-mannosides. Here, we characterized β-1,2-mannobiose phosphorylase from Listeria innocua (Lin0857) and determined its crystal structures complexed with β-1,2-linked mannooligosaccharides. β-1,2-Mannotriose was bound in a U-shape, interacting with a phosphate analog at both ends. Lin0857 has a unique dimer structure connected by a loop, and a significant open-close loop displacement was observed for substrate entry. A long loop, which is exclusively present in Lin0857, covers the active site to limit the pocket size. A structural basis for substrate recognition and phosphorolysis was provided.
Organizational Affiliation:
Department of Biotechnology, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.