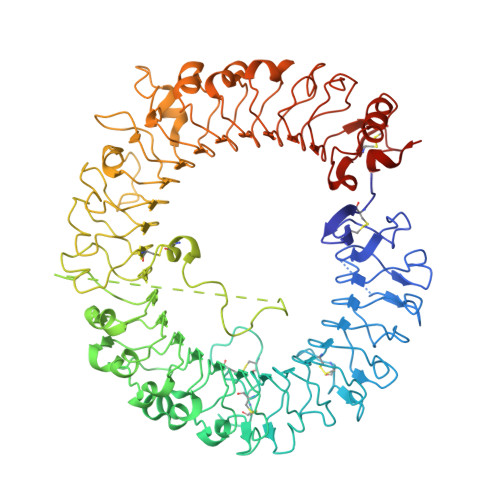
Small-molecule inhibition of TLR8 through stabilization of its resting state
Zhang, S., Hu, Z., Tanji, H., Jiang, S., Das, N., Li, J., Sakaniwa, K., Jin, J., Bian, Y., Ohto, U., Shimizu, T., Yin, H.(2018) Nat Chem Biol 14: 58-64
- PubMed: 29155428
- DOI: https://doi.org/10.1038/nchembio.2518
- Primary Citation of Related Structures:
5WYX, 5WYZ - PubMed Abstract:
Endosomal Toll-like receptors (TLR3, TLR7, TLR8, and TLR9) are highly analogous sensors for various viral or bacterial RNA and DNA molecular patterns. Nonetheless, few small molecules can selectively modulate these TLRs. In this manuscript, we identified the first human TLR8-specific small-molecule antagonists via a novel inhibition mechanism. Crystal structures of two distinct TLR8-ligand complexes validated a unique binding site on the protein-protein interface of the TLR8 homodimer. Upon binding to this new site, the small-molecule ligands stabilize the preformed TLR8 dimer in its resting state, preventing activation. As a proof of concept of their therapeutic potential, we have demonstrated that these drug-like inhibitors are able to suppress TLR8-mediated proinflammatory signaling in various cell lines, human primary cells, and patient specimens. These results not only suggest a novel strategy for TLR inhibitor design, but also shed critical mechanistic insight into these clinically important immune receptors.
Organizational Affiliation:
School of Pharmaceutical Sciences, Center of Basic Molecular Science, Key Laboratory of Bioorganic Phosphorus Chemistry and Chemical Biology (Ministry of Education), Department of Chemistry, Tsinghua University, Beijing, China.