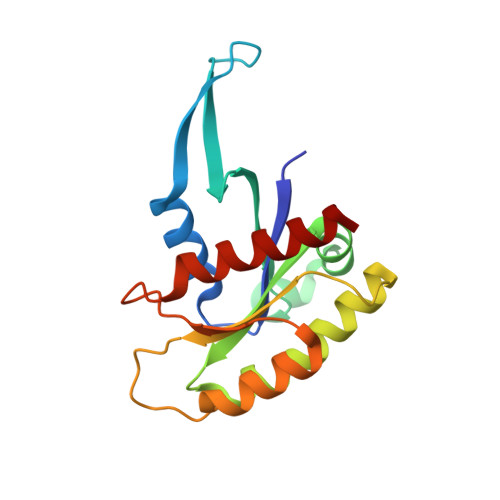
Structures of N-terminally processed KRAS provide insight into the role of N-acetylation.
Dharmaiah, S., Tran, T.H., Messing, S., Agamasu, C., Gillette, W.K., Yan, W., Waybright, T., Alexander, P., Esposito, D., Nissley, D.V., McCormick, F., Stephen, A.G., Simanshu, D.K.(2019) Sci Rep 9: 10512-10512
- PubMed: 31324887
- DOI: https://doi.org/10.1038/s41598-019-46846-w
- Primary Citation of Related Structures:
6M9W, 6MBQ, 6MBT, 6MBU, 6P0Z - PubMed Abstract:
Although post-translational modification of the C-terminus of RAS has been studied extensively, little is known about N-terminal processing. Mass spectrometric characterization of KRAS expressed in mammalian cells showed cleavage of the initiator methionine (iMet) and N-acetylation of the nascent N-terminus. Interestingly, structural studies on GDP- and GMPPNP-bound KRAS lacking the iMet and N-acetylation resulted in Mg 2+ -free structures of KRAS with flexible N-termini. In the Mg 2+ -free KRAS-GDP structure, the flexible N-terminus causes conformational changes in the interswitch region resulting in a fully open conformation of switch I. In the Mg 2+ -free KRAS-GMPPNP structure, the flexible N-terminus causes conformational changes around residue A59 resulting in the loss of Mg 2+ and switch I in the inactive state 1 conformation. Structural studies on N-acetylated KRAS-GDP lacking the iMet revealed the presence of Mg 2+ and a conformation of switch regions also observed in the structure of GDP-bound unprocessed KRAS with the iMet. In the absence of the iMet, the N-acetyl group interacts with the central beta-sheet and stabilizes the N-terminus and the switch regions. These results suggest there is crosstalk between the N-terminus and the Mg 2+ binding site, and that N-acetylation plays an important role by stabilizing the N-terminus of RAS upon excision of the iMet.
Organizational Affiliation:
NCI RAS Initiative, Cancer Research Technology Program, Frederick National Laboratory for Cancer Research, Leidos Biomedical Research, Inc., Frederick, MD, 21701, USA.