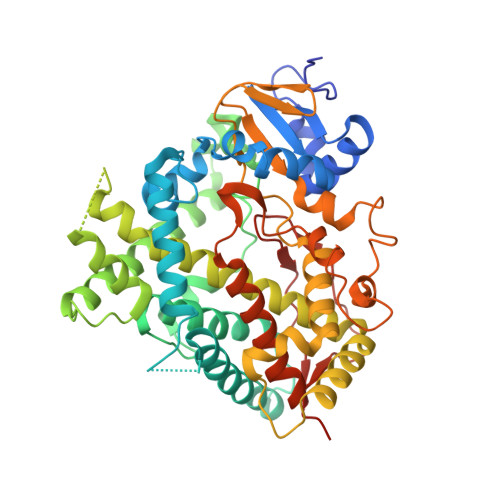
Human Cytochrome P450 1A1 Adapts Active Site for Atypical Nonplanar Substrate.
Bart, A.G., Takahashi, R.H., Wang, X., Scott, E.E.(2020) Drug Metab Dispos 48: 86-92
- PubMed: 31757797
- DOI: https://doi.org/10.1124/dmd.119.089607
- Primary Citation of Related Structures:
6O5Y - PubMed Abstract:
The human cytochrome P450 1A1 (CYP1A1) is well known for chemical activation of procarcinogens and often has a substrate scope of small and highly planar compounds. Substrates deviating from these characteristics are certainly known, but how these larger and nonplanar substrates are accommodated and oriented within the CYP1A1 active site is not understood. Herein a new X-ray structure of CYP1A1 bound to the pan-Pim kinase inhibitor GDC-0339 reveals how the CYP1A1 active site cavity is reconfigured to bind larger and nonplanar compounds. The shape and size of the cavity are controlled by structural elements in the active site roof, with major changes in the conformation of the F helix break and relocation of Phe224 from the active site to the protein surface. This altered CYP1A1 active site architecture is consistent with the proposed mechanism for CYP1A1 generation of an unusual aminoazepane-rearranged metabolite for this substrate. SIGNIFICANCE STATEMENT: Cytochrome P450 1A1 metabolizes drugs, procarcinogens, and toxins and although previous structures have revealed how its stereotypical planar, aromatic compounds are accommodated in the CYP1A1 active site, this is not the case for flexible and nonplanar compounds. The current work determines the X-ray structure of CYP1A1 with such a flexible, nonplanar Pim kinase inhibitor, revealing significant modification of the CYP1A1 roof that accommodate this preclinical candidate and support an unusual intramolecular rearrangement reaction.
Organizational Affiliation:
Program in Biophysics (A.G.B., E.E.S.) and Departments of Medicinal Chemistry and Pharmacology (E.E.S.), University of Michigan, Ann Arbor, Michigan; and Department of Drug Metabolism and Pharmacokinetics (R.H.T.) and Department of Discovery Chemistry (X.W.), Genentech Inc., South San Francisco, California.