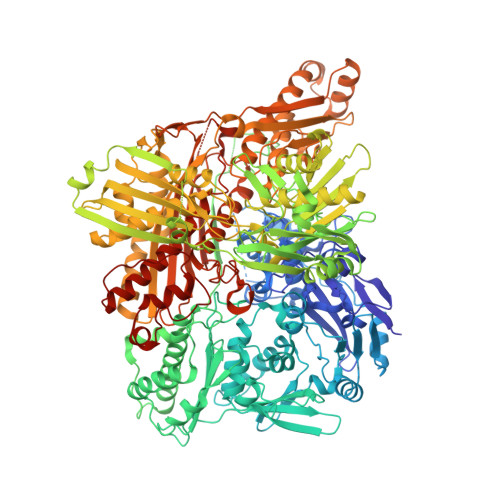
Human aldehyde oxidase (hAOX1): structure determination of the Moco-free form of the natural variant G1269R and biophysical studies of single nucleotide polymorphisms.
Mota, C., Esmaeeli, M., Coelho, C., Santos-Silva, T., Wolff, M., Foti, A., Leimkuhler, S., Romao, M.J.(2019) FEBS Open Bio 9: 925-934
- PubMed: 30985987
- DOI: https://doi.org/10.1002/2211-5463.12617
- Primary Citation of Related Structures:
6Q6Q - PubMed Abstract:
Human aldehyde oxidase (hAOX1) is a molybdenum enzyme with high toxicological importance, but its physiological role is still unknown. hAOX1 metabolizes different classes of xenobiotics and is one of the main drug-metabolizing enzymes in the liver, along with cytochrome P450. hAOX1 oxidizes and inactivates a large number of drug molecules and has been responsible for the failure of several phase I clinical trials. The interindividual variability of drug-metabolizing enzymes caused by single nucleotide polymorphisms (SNPs) is highly relevant in pharmaceutical treatments. In this study, we present the crystal structure of the inactive variant G1269R, revealing the first structure of a molybdenum cofactor (Moco)-free form of hAOX1. These data allowed to model, for the first time, the flexible Gate 1 that controls access to the active site. Furthermore, we inspected the thermostability of wild-type hAOX1 and hAOX1 with various SNPs (L438V, R1231H, G1269R or S1271L) by CD spectroscopy and ThermoFAD, revealing that amino acid exchanges close to the Moco site can impact protein stability up to 10 °C. These results correlated with biochemical and structural data and enhance our understanding of hAOX1 and the effect of SNPs in the gene encoding this enzyme in the human population. ENZYMES: Aldehyde oxidase (EC1.2.3.1); xanthine dehydrogenase (EC1.17.1.4); xanthine oxidase (EC1.1.3.2). DATABASES: Structural data are available in the Protein Data Bank under the accession number 6Q6Q.
Organizational Affiliation:
UCIBIO, Departamento de Química, Faculdade de Ciências e Tecnologia, Universidade Nova de Lisboa, Caparica, Portugal.