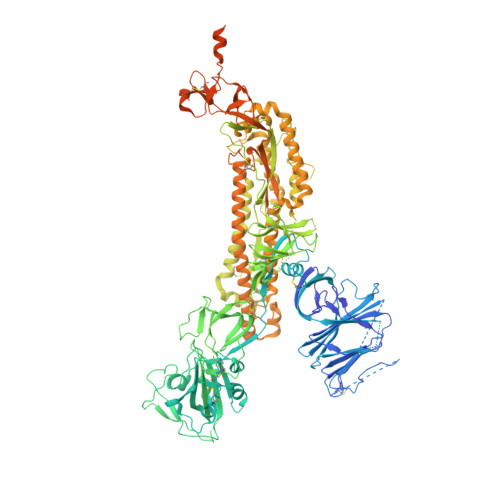
D614G Mutation Alters SARS-CoV-2 Spike Conformation and Enhances Protease Cleavage at the S1/S2 Junction.
Gobeil, S.M., Janowska, K., McDowell, S., Mansouri, K., Parks, R., Manne, K., Stalls, V., Kopp, M.F., Henderson, R., Edwards, R.J., Haynes, B.F., Acharya, P.(2021) Cell Rep 34: 108630-108630
- PubMed: 33417835
- DOI: https://doi.org/10.1016/j.celrep.2020.108630
- PubMed Abstract:
The severe acute respiratory coronavirus 2 (SARS-CoV-2) spike (S) protein is the target of vaccine design efforts to end the coronavirus disease 2019 (COVID-19) pandemic. Despite a low mutation rate, isolates with the D614G substitution in the S protein appeared early during the pandemic and are now the dominant form worldwide. Here, we explore S conformational changes and the effects of the D614G mutation on a soluble S ectodomain construct. Cryoelectron microscopy (cryo-EM) structures reveal altered receptor binding domain (RBD) disposition; antigenicity and proteolysis experiments reveal structural changes and enhanced furin cleavage efficiency of the G614 variant. Furthermore, furin cleavage alters the up/down ratio of the RBDs in the G614 S ectodomain, demonstrating an allosteric effect on RBD positioning triggered by changes in the SD2 region, which harbors residue 614 and the furin cleavage site. Our results elucidate SARS-CoV-2 S conformational landscape and allostery and have implications for vaccine design.
Organizational Affiliation:
Duke Human Vaccine Institute, Durham, NC 27710, USA.