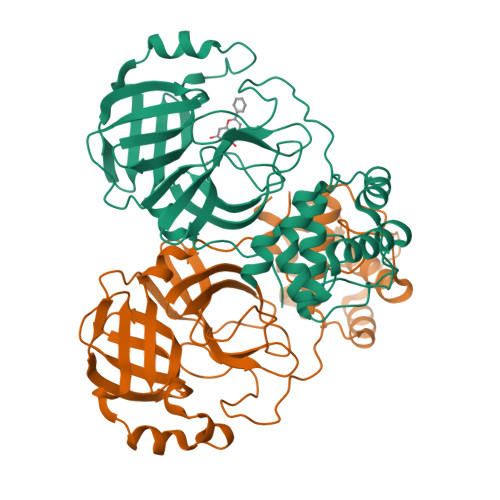
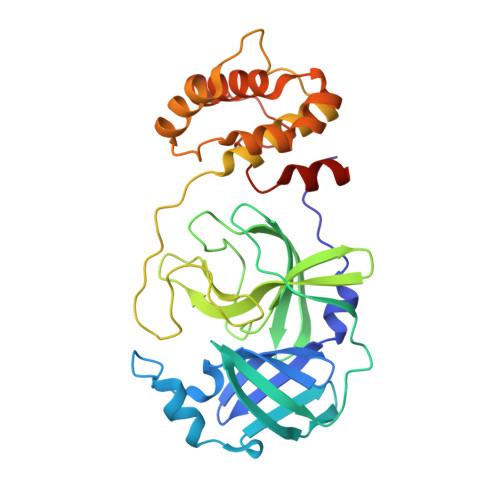
Crystal structure of SARS-CoV 3C-like protease with baicalein.
Feng, J., Li, D., Zhang, J., Yin, X., Li, J.(2022) Biochem Biophys Res Commun 611: 190-194
- PubMed: 35490659
- DOI: https://doi.org/10.1016/j.bbrc.2022.04.086
- Primary Citation of Related Structures:
7XAX - PubMed Abstract:
The 3C-like protease (M pro , 3CL pro ) plays a key role in the replication process in coronaviruses (CoVs). The M pro is an essential enzyme mediates CoVs replication and is a promising target for development of antiviral drugs. Until now, baicalein has been shown the specific activity for SARS-CoV M pro in vitro experiments. In this study, we resolved the SARS-CoV M pro with baicalein by X-ray diffraction at 2.25 Å (PDB code 7XAX), which provided a structural basis for the research and development of baicalein as an anti-CoVs drug.
Organizational Affiliation:
Applied Biology Laboratory, College of Pharmaceutical and Biological Engineering, Shenyang University of Chemical Technology, Shenyang, 110142, China.