A multi-reservoir extruder for time-resolved serial protein crystallography and compound screening at X-ray free-electron lasers.
Wranik, M., Kepa, M.W., Beale, E.V., James, D., Bertrand, Q., Weinert, T., Furrer, A., Glover, H., Gashi, D., Carrillo, M., Kondo, Y., Stipp, R.T., Khusainov, G., Nass, K., Ozerov, D., Cirelli, C., Johnson, P.J.M., Dworkowski, F., Beale, J.H., Stubbs, S., Zamofing, T., Schneider, M., Krauskopf, K., Gao, L., Thorn-Seshold, O., Bostedt, C., Bacellar, C., Steinmetz, M.O., Milne, C., Standfuss, J.(2023) Nat Commun 14: 7956-7956
- PubMed: 38042952
- DOI: https://doi.org/10.1038/s41467-023-43523-5
- Primary Citation of Related Structures:
8CL5, 8CL6, 8CL7, 8CL8, 8CL9, 8CLB, 8CLC, 8CLD, 8CLE, 8CLF, 8CLG, 8CLH - PubMed Abstract:
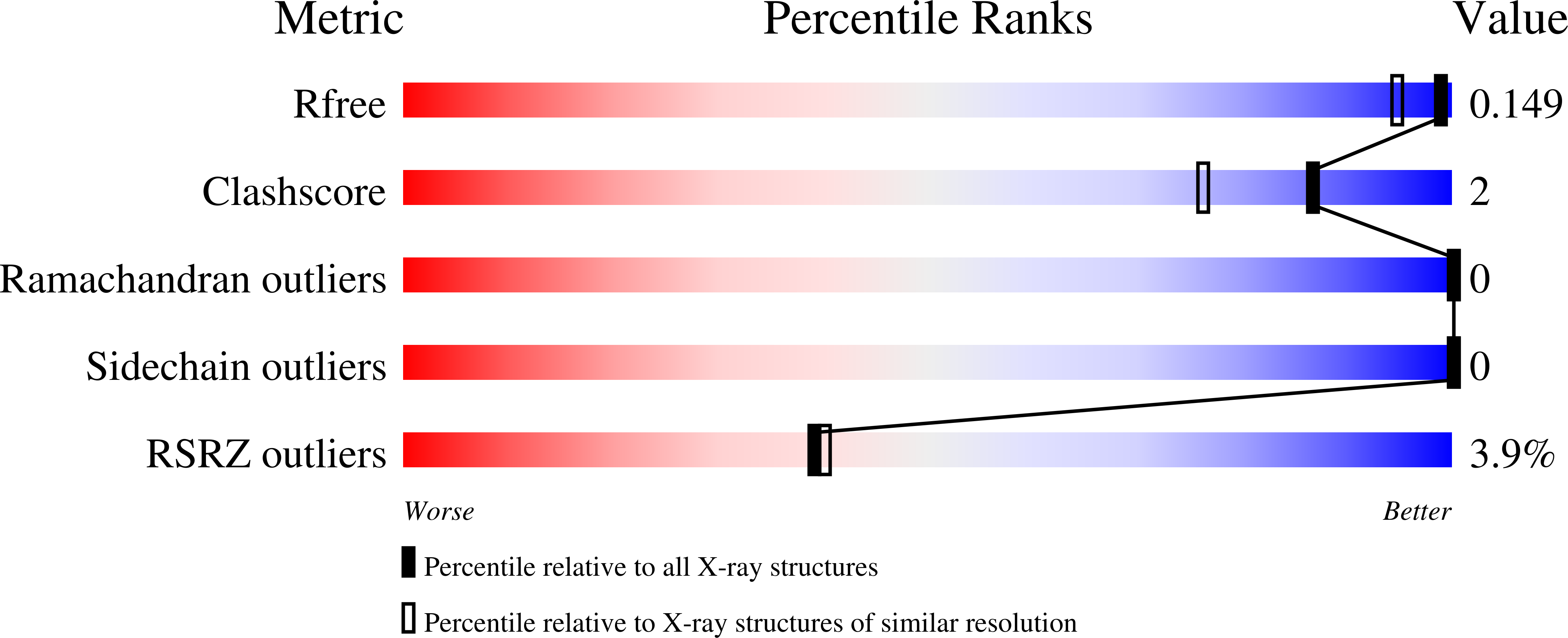
Serial crystallography at X-ray free-electron lasers (XFELs) permits the determination of radiation-damage free static as well as time-resolved protein structures at room temperature. Efficient sample delivery is a key factor for such experiments. Here, we describe a multi-reservoir, high viscosity extruder as a step towards automation of sample delivery at XFELs. Compared to a standard single extruder, sample exchange time was halved and the workload of users was greatly reduced. In-built temperature control of samples facilitated optimal extrusion and supported sample stability. After commissioning the device with lysozyme crystals, we collected time-resolved data using crystals of a membrane-bound, light-driven sodium pump. Static data were also collected from the soluble protein tubulin that was soaked with a series of small molecule drugs. Using these data, we identify low occupancy (as little as 30%) ligands using a minimal amount of data from a serial crystallography experiment, a result that could be exploited for structure-based drug design.
Organizational Affiliation:
Laboratory of Biomolecular Research, Division of Biology and Chemistry, Paul Scherrer Institut, Villigen-PSI, Villigen, 5232, Switzerland. maximilian.wranik@psi.ch.