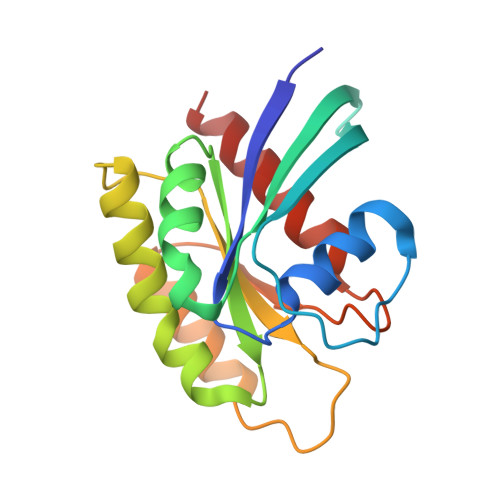
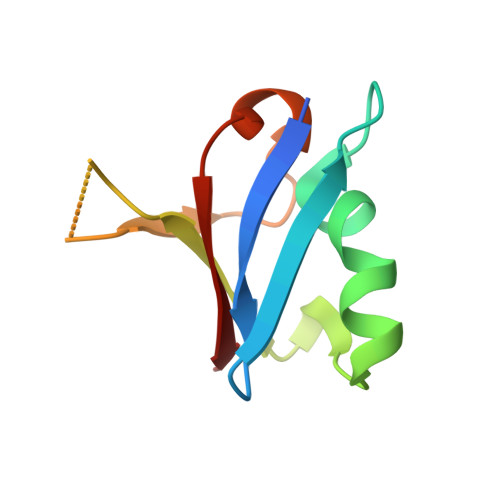
Reduced dynamic complexity allows structure elucidation of an excited state of KRAS G13D .
Chao, F.A., Chan, A.H., Dharmaiah, S., Schwieters, C.D., Tran, T.H., Taylor, T., Ramakrishnan, N., Esposito, D., Nissley, D.V., McCormick, F., Simanshu, D.K., Cornilescu, G.(2023) Commun Biol 6: 594-594
- PubMed: 37268708
- DOI: https://doi.org/10.1038/s42003-023-04960-6
- Primary Citation of Related Structures:
8EBZ, 8EPW - PubMed Abstract:
Localized dynamics of RAS, including regions distal to the nucleotide-binding site, is of high interest for elucidating the mechanisms by which RAS proteins interact with effectors and regulators and for designing inhibitors. Among several oncogenic mutants, methyl relaxation dispersion experiments reveal highly synchronized conformational dynamics in the active (GMPPNP-bound) KRAS G13D , which suggests an exchange between two conformational states in solution. Methyl and 31 P NMR spectra of active KRAS G13D in solution confirm a two-state ensemble interconverting on the millisecond timescale, with a major P γ atom peak corresponding to the dominant State 1 conformation and a secondary peak indicating an intermediate state different from the known State 2 conformation recognized by RAS effectors. High-resolution crystal structures of active KRAS G13D and KRAS G13D -RAF1 RBD complex provide snapshots of the State 1 and 2 conformations, respectively. We use residual dipolar couplings to solve and cross-validate the structure of the intermediate state of active KRAS G13D , showing a conformation distinct from those of States 1 and 2 outside the known flexible switch regions. The dynamic coupling between the conformational exchange in the effector lobe and the breathing motion in the allosteric lobe is further validated by a secondary mutation in the allosteric lobe, which affects the conformational population equilibrium.
Organizational Affiliation:
NCI RAS Initiative, Cancer Research Technology Program, Frederick National Laboratory for Cancer Research, Leidos Biomedical Research, Frederick, MD, 21701, USA. fa-an.chao@nih.gov.