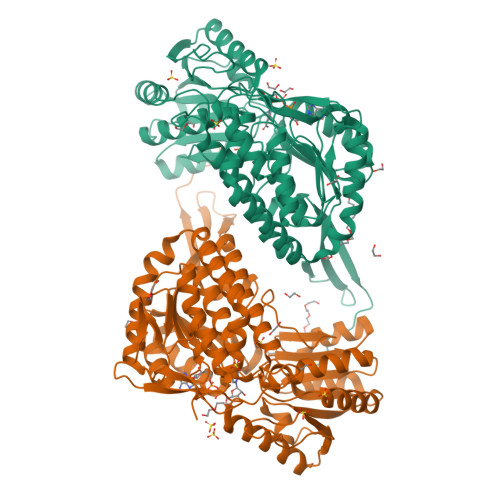
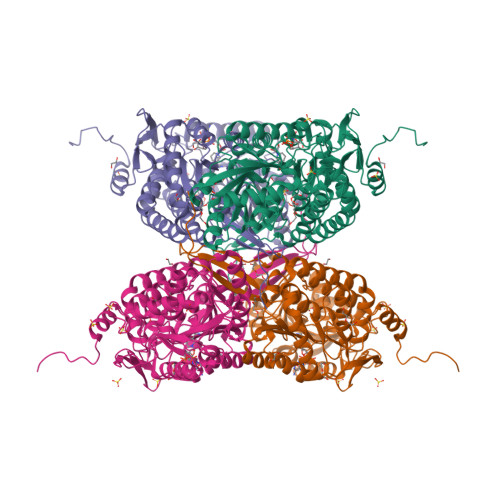
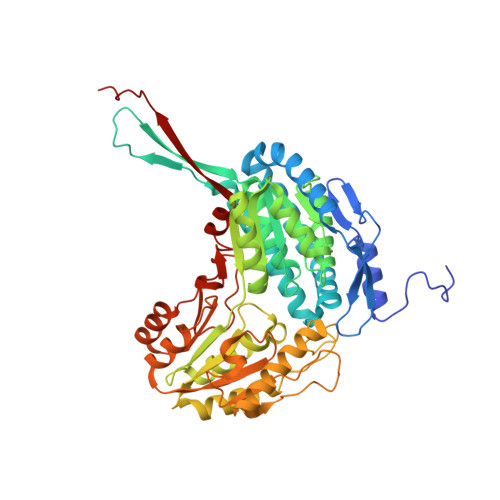
A study on abiotic stress responses of aldehyde dehydrogenase (ALDH) superfamilies in moss and barley focused on members linked to the GABA shunt pathway
Kopecny, D.J., Belicek, J., Vigouroux, A., Luptakova, E., Koncitikova, R., von Schwartzenberg, K., Cavar Zeljkovic, S., Supikova, k., Gruz, J., Valarik, m., Bergougnoux, V., Kopecny, D., Morera, S., Kopecna, M.To be published.