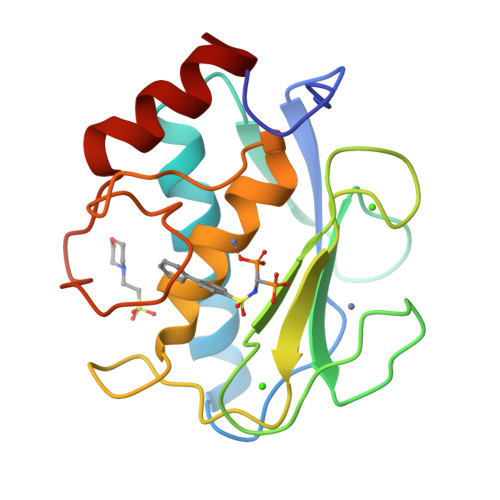
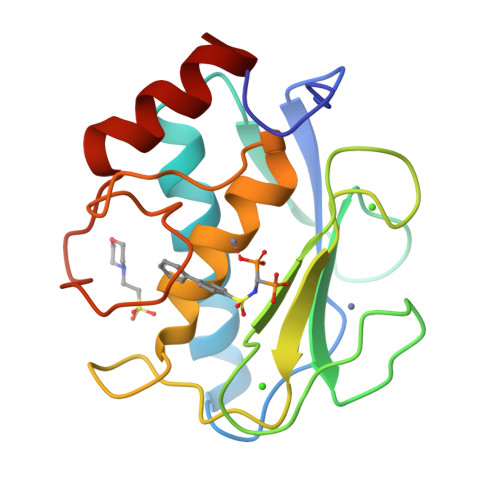
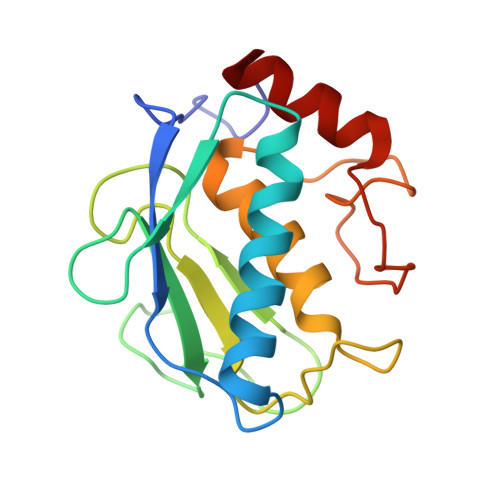
Bone-Seeking Matrix Metalloproteinase Inhibitors for the Treatment of Skeletal Malignancy.
Laghezza, A., Piemontese, L., Brunetti, L., Caradonna, A., Agamennone, M., Di Pizio, A., Pochetti, G., Montanari, R., Capelli, D., Tauro, M., Loiodice, F., Tortorella, P.(2020) Pharmaceuticals (Basel) 13
- PubMed: 32492898
- DOI: https://doi.org/10.3390/ph13060113
- Primary Citation of Related Structures:
4QKZ - PubMed Abstract:
Matrix metalloproteinases (MMPs) are a family of enzymes involved at different stages of cancer progression and metastasis. We previously identified a novel class of bisphosphonic inhibitors, selective for MMPs crucial for bone remodeling, such as MMP-2. Due to the increasing relevance of specific MMPs at various stages of tumor malignancy, we focused on improving potency towards certain isoforms. Here, we tackled MMP-9 because of its confirmed role in tumor invasion, metastasis, angiogenesis, and immuno-response, making it an ideal target for cancer therapy. Using a computational analysis, we designed and characterized potent MMP-2/MMP-9 inhibitors. This is a promising approach to develop and clinically translate inhibitors that could be used in combination with standard care therapy for the treatment of skeletal malignancies.
Organizational Affiliation:
Department of Pharmacy and Pharmaceutical Sciences, University of Bari "A. Moro", via E. Orabona 4, 70125 Bari, Italy.