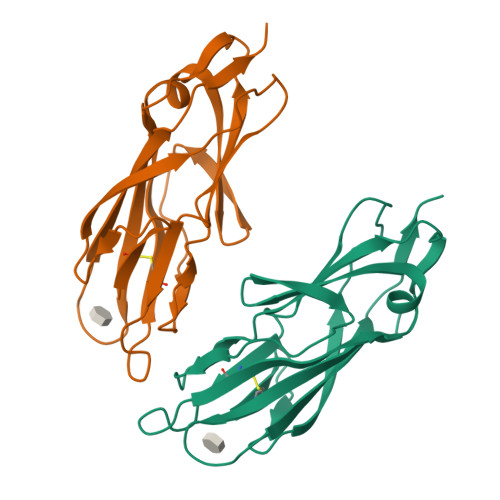
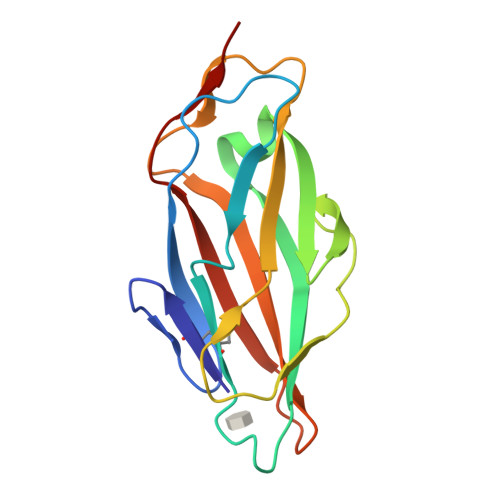
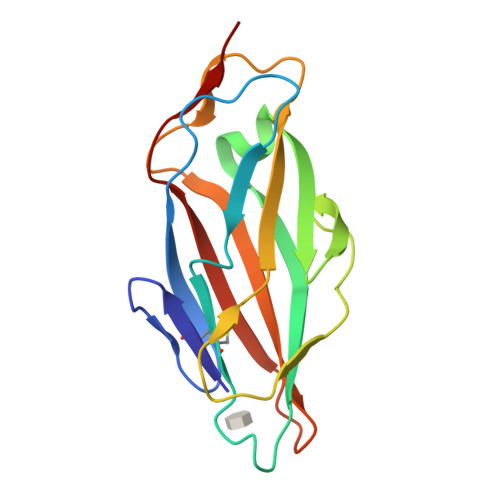
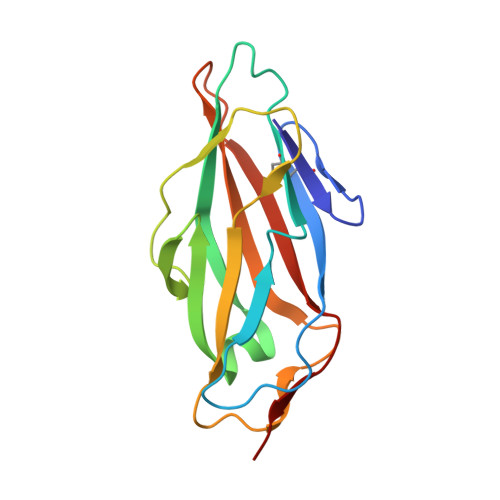
High-Affinity Carbohydrate-Lectin Interactions: How Nature Makes it Possible
Zihlmann, P., Jiang, X., Sager, C.P., Fiege, B., Jakob, R.P., Siegrist, S., Zalewski, A., Rabbani, S., Eris, D., Silbermann, M., Pang, L., Muhlethaler, T., Sharpe, T., Maier, T., Ernst, B.To be published.