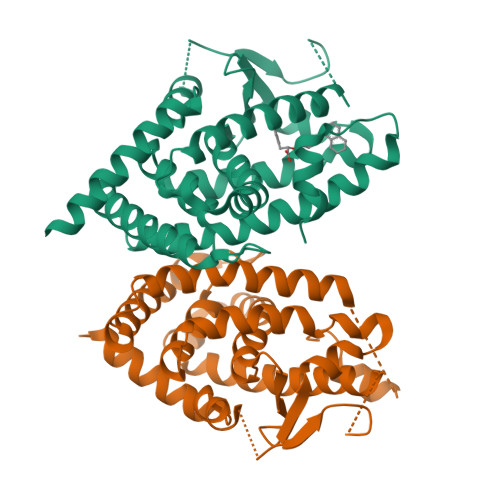
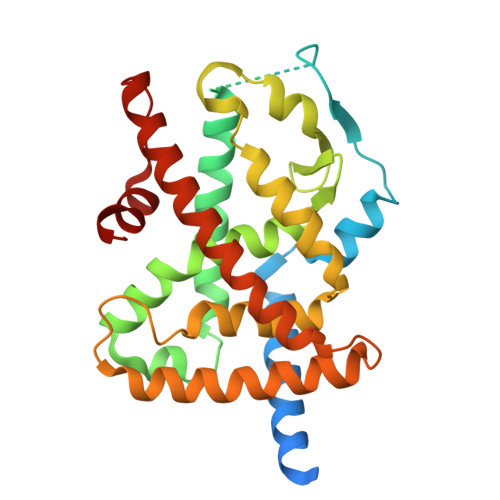
A chemical modification of a peroxisome proliferator-activated receptor pan agonist produced a shift to a new dual alpha/gamma partial agonist endowed with mitochondrial pyruvate carrier inhibition and antidiabetic properties.
Laghezza, A., Cerchia, C., Genovese, M., Montanari, R., Capelli, D., Wackerlig, J., Simic, S., Falbo, E., Pecora, L., Leuci, R., Brunetti, L., Piemontese, L., Tortorella, P., Biswas, A., Singh, R.P., Tambe, S., Sudeep, C.A., Pattnaik, A.K., Jayaprakash, V., Paoli, P., Lavecchia, A., Loiodice, F.(2024) Eur J Med Chem 275: 116567-116567
- PubMed: 38865743
- DOI: https://doi.org/10.1016/j.ejmech.2024.116567
- Primary Citation of Related Structures:
8RCE, 8REJ - PubMed Abstract:
New analogs of the PPAR pan agonist AL29-26 encompassed ligand (S)-7 showing potent activation of PPARα and -γ subtypes as a partial agonist. In vitro experiments and docking studies in the presence of PPAR antagonists were performed to help interpretation of biological data and investigate the main interactions at the binding sites. Further in vitro experiments showed that (S)-7 induced anti-steatotic effects and enhancement of the glucose uptake. This latter effect could be partially ascribed to a significant inhibition of the mitochondrial pyruvate carrier demonstrating that (S)-7 also acted through insulin-independent mechanisms. In vivo experiments showed that this compound reduced blood glucose and lipid levels in a diabetic mice model displaying no toxicity on bone, kidney, and liver. To our knowledge, this is the first example of dual PPARα/γ partial agonist showing these combined effects representing, therefore, the potential lead of new drugs for treatment of dyslipidemic type 2 diabetes.
Organizational Affiliation:
Dipartimento Farmacia-Scienze del Farmaco, Università degli Studi di Bari "Aldo Moro", via Orabona 4, 70125, Bari, Italy.