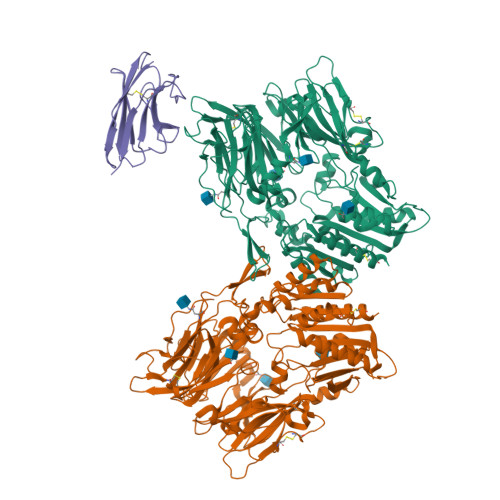
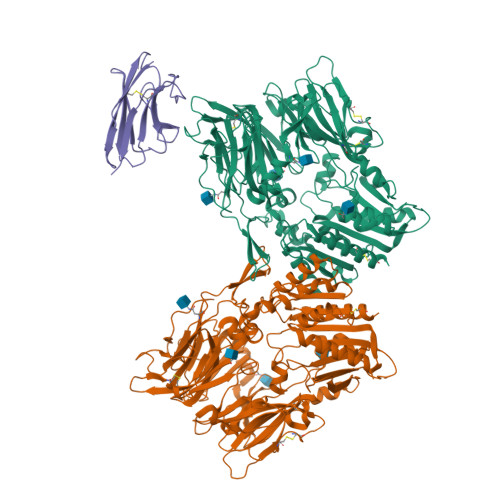
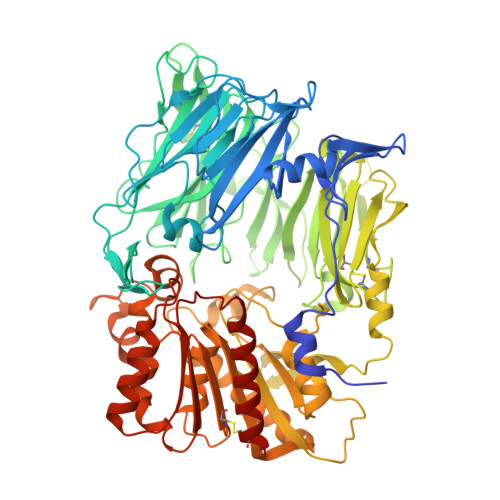
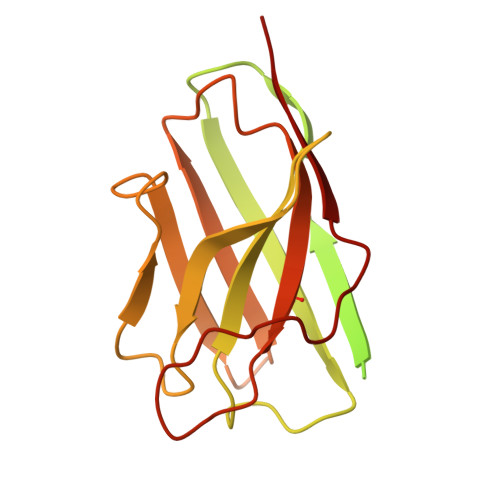
Cryo-electron microscopy reveals a single domain antibody with a unique binding epitope on fibroblast activation protein alpha.
Xu, Z., Sinha, A., Pandya, D.N., Schnicker, N.J., Wadas, T.J.(2025) RSC Chem Biol
- PubMed: 39975582
- DOI: https://doi.org/10.1039/d4cb00267a
- Primary Citation of Related Structures:
9DVQ, 9DVR - PubMed Abstract:
Fibroblast activation protein alpha (FAP) is a serine protease that is expressed at basal levels in benign tissues but is overexpressed in a variety of pathologies, including cancer. Despite this unique expression profile, designing functional diagnostic and therapeutic agents that effectively target this biomarker remains elusive. Here we report the structural characterization of the interaction between a novel single domain antibody (sdAb), I3, and FAP using cryo-electron microscopy. The reconstructions were determined to a resolution of 2.7 Å and contained two distinct populations; one I3 bound and two I3 molecules bound to the FAP dimer. In both cases, the sdAb bound a unique epitope that was distinct from the active site of the enzyme. Furthermore, this report describes the rational mutation of specific residues within the complementarity determining region 3 (CDR3) loop to enhance affinity and selectivity of the I3 molecule for FAP. This report represents the first sdAb-FAP structure to be described in the literature.
Organizational Affiliation:
Protein and Crystallography Facility, University of Iowa Iowa City Iowa 52242 USA nicholas-schnicker@uiowa.edu.