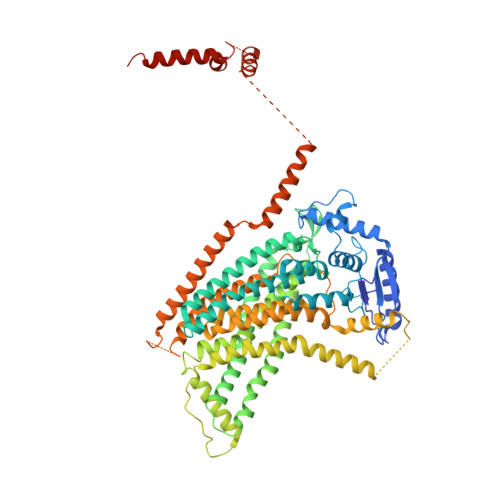
Structural basis of closed groove scrambling by a TMEM16 protein.
Feng, Z., Alvarenga, O.E., Accardi, A.(2024) Nat Struct Mol Biol
- PubMed: 38684930
- DOI: https://doi.org/10.1038/s41594-024-01284-9
- Primary Citation of Related Structures:
8TOI, 8TOK, 8TOL, 8TPM, 8TPN, 8TPO, 8TPP, 8TPQ, 8TPR, 8TPS, 8TPT - PubMed Abstract:
Activation of Ca 2+ -dependent TMEM16 scramblases induces phosphatidylserine externalization, a key step in multiple signaling processes. Current models suggest that the TMEM16s scramble lipids by deforming the membrane near a hydrophilic groove and that Ca 2+ dependence arises from the different association of lipids with an open or closed groove. However, the molecular rearrangements underlying groove opening and how lipids reorganize outside the closed groove remain unknown. Here we directly visualize how lipids associate at the closed groove of Ca 2+ -bound fungal nhTMEM16 in nanodiscs using cryo-EM. Functional experiments pinpoint lipid-protein interaction sites critical for closed groove scrambling. Structural and functional analyses suggest groove opening entails the sequential appearance of two π-helical turns in the groove-lining TM6 helix and identify critical rearrangements. Finally, we show that the choice of scaffold protein and lipids affects the conformations of nhTMEM16 and their distribution, highlighting a key role of these factors in cryo-EM structure determination.
Organizational Affiliation:
Department of Anesthesiology, Weill Cornell Medical College, New York, NY, USA.