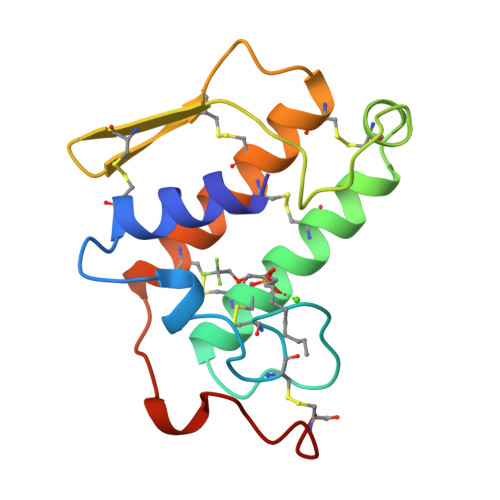
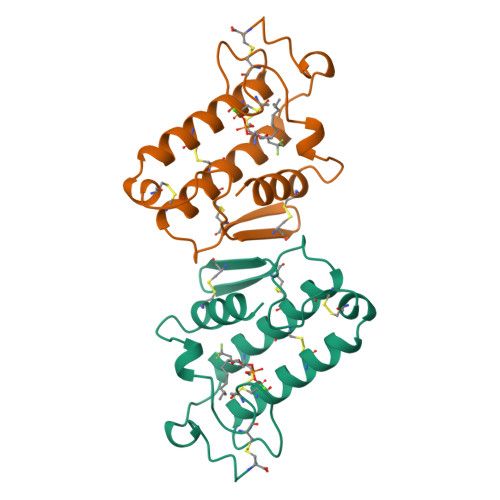
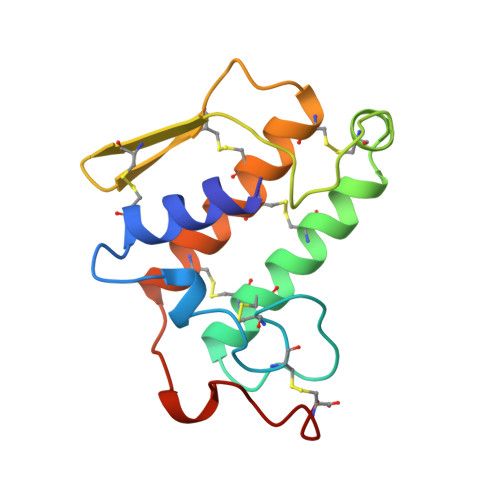
Crystal structure of the complex of bovine pancreatic phospholipase A2 with the inhibitor 1-hexadecyl-3-(trifluoroethyl)-sn-glycero-2-phosphomethanol,.
Sekar, K., Eswaramoorthy, S., Jain, M.K., Sundaralingam, M.(1997) Biochemistry 36: 14186-14191
- PubMed: 9369492
- DOI: https://doi.org/10.1021/bi971370b
- Primary Citation of Related Structures:
1FDK - PubMed Abstract:
The structure of recombinant bovine pancreatic phospholipase A2 (PLA2) complexed with the competitive inhibitor 1-hexadecyl-3-(trifluoroethyl)-sn-glycero -2-phosphomethanol (hereafter MJ33), a phospholipid analogue without the sn-3 phosphodiester group, has been determined. The crystals are trigonal, space group P3121, a = b = 46.36 A and c = 102.56 A, and isomorphous to the recombinant PLA2 with one molecule in the asymmetric unit. The structure was refined using 8082 reflections between 8.0 and 1.91 A resolution to a final R-value of 18.4% [Rfree = 28.0%]. The model includes 957 protein atoms, 86 water molecules, one calcium ion, and 26 non-hydrogen atoms of the inhibitor MJ33. The overall tertiary fold of the complex is very similar to that of the inhibitor-free recombinant PLA2 with a root mean square deviation of 0.32 A for all the backbone atoms. The electron density of the surface loop residues 62-66 is clear and ordered, unlike the other trigonal bovine PLA2 structures done to date. This structural change could be responsible for the interfacial allosteric activation, which thermodynamically relates the enhanced binding of the substrate mimic to the active site of the enzyme. MJ33 is tightly bound in the active-site cleft, dislodging the equatorial coordinated calcium water (W5), the putative catalytic water W6, and the neighboring water W7. The axial coordinated calcium water is missing; thus the hexacoordinated calcium is a monocapped pentagonal pyramid. Although MJ33 is a sn-2 tetrahedral mimic, its phosphate binds to PLA2 differently from the sn-2 phosphonate analogue of phospholipids, another tetrahedral mimic. The knowledge of the active-site geometry of MJ33 would be useful in the design of more useful therapeutic agents for PLA2.
Organizational Affiliation:
Biological Macromolecular Structure Center, Department of Chemistry, Ohio State University, Columbus, Ohio 43210, USA.