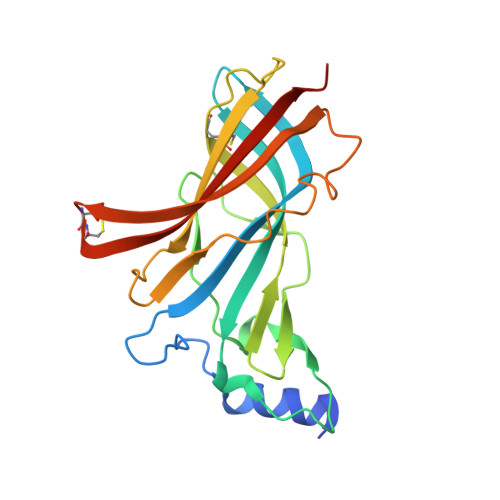
Crystal structure of an ACh-binding protein reveals the ligand-binding domain of nicotinic receptors.
Brejc, K., van Dijk, W.J., Klaassen, R.V., Schuurmans, M., van Der Oost, J., Smit, A.B., Sixma, T.K.(2001) Nature 411: 269-276
- PubMed: 11357122
- DOI: https://doi.org/10.1038/35077011
- Primary Citation of Related Structures:
1I9B - PubMed Abstract:
Pentameric ligand gated ion-channels, or Cys-loop receptors, mediate rapid chemical transmission of signals. This superfamily of allosteric transmembrane proteins includes the nicotinic acetylcholine (nAChR), serotonin 5-HT3, gamma-aminobutyric-acid (GABAA and GABAC) and glycine receptors. Biochemical and electrophysiological information on the prototypic nAChRs is abundant but structural data at atomic resolution have been missing. Here we present the crystal structure of molluscan acetylcholine-binding protein (AChBP), a structural and functional homologue of the amino-terminal ligand-binding domain of an nAChR alpha-subunit. In the AChBP homopentamer, the protomers have an immunoglobulin-like topology. Ligand-binding sites are located at each of five subunit interfaces and contain residues contributed by biochemically determined 'loops' A to F. The subunit interfaces are highly variable within the ion-channel family, whereas the conserved residues stabilize the protomer fold. This AChBP structure is relevant for the development of drugs against, for example, Alzheimer's disease and nicotine addiction.
Organizational Affiliation:
Division of Molecular Carcinogenesis, Netherlands Cancer Institute, Plesmanlaan 121, 1066 CX Amsterdam, The Netherlands.