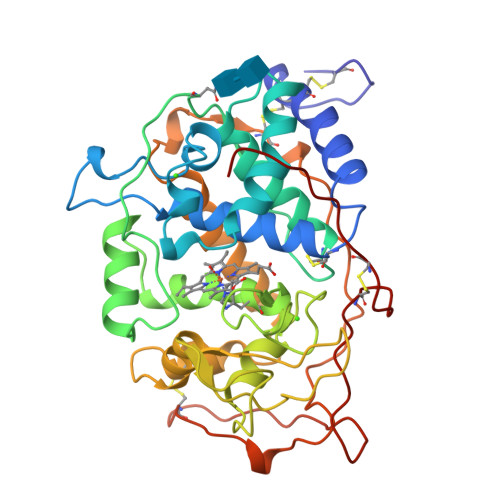
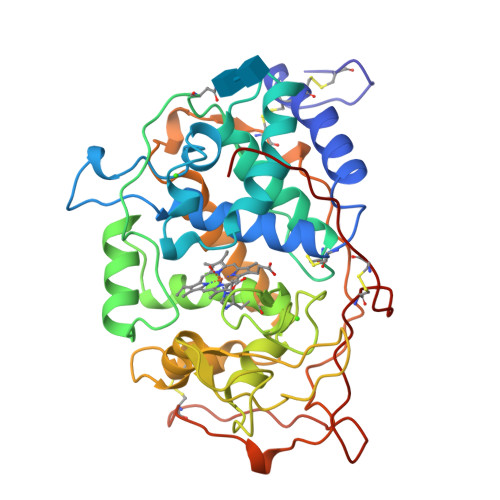
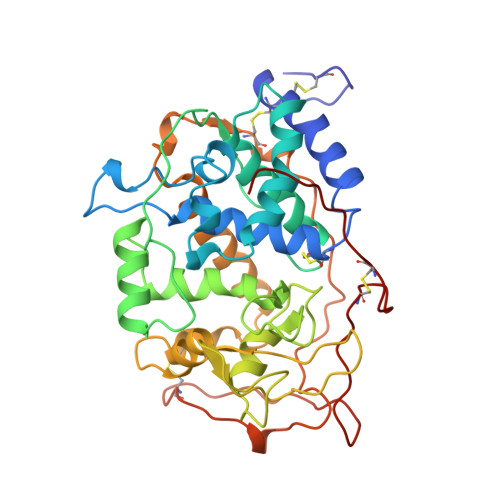
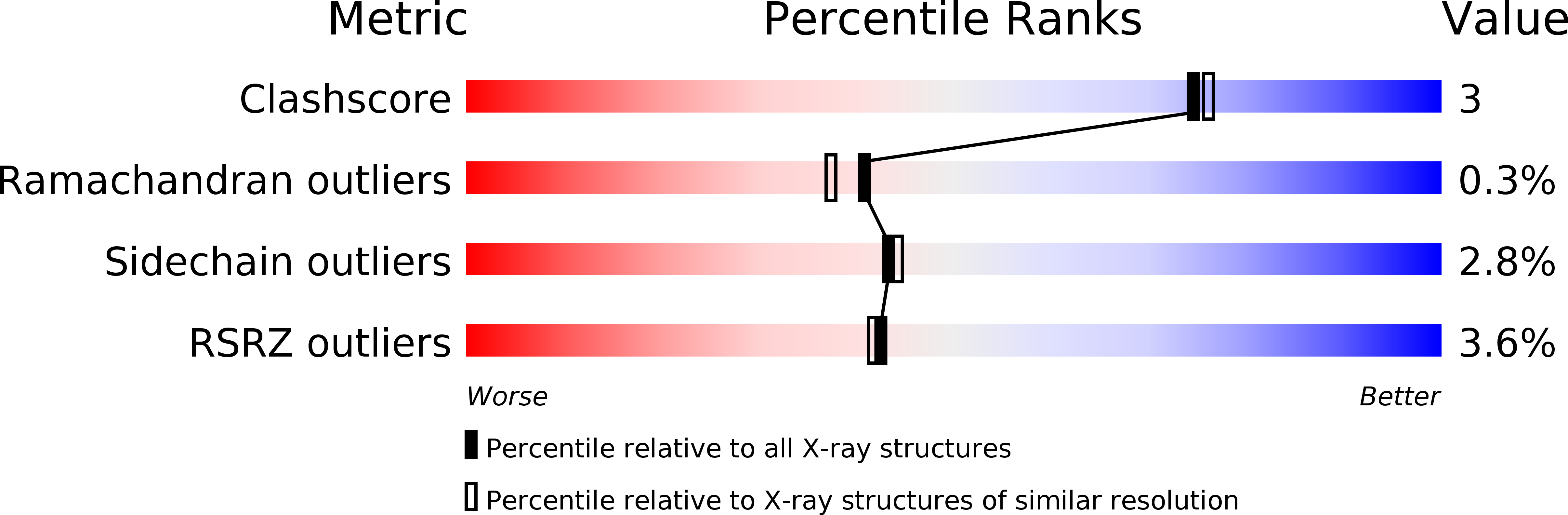Crystal structures of substrate binding site mutants of manganese peroxidase.
Sundaramoorthy, M., Kishi, K., Gold, M.H., Poulos, T.L.(1997) J Biological Chem 272: 17574-17580
- PubMed: 9211904
- DOI: https://doi.org/10.1074/jbc.272.28.17574
- Primary Citation of Related Structures:
1MN1, 1MN2 - PubMed Abstract:
Manganese peroxidase (MnP), an extracellular heme enzyme from the lignin-degrading basidiomycetous fungus, Phanerochaete chrysosporium, catalyzes the oxidation of MnII to MnIII. The latter, acting as a diffusible redox mediator, is capable of oxidizing a variety of lignin model compounds. The proposed MnII binding site of MnP consists of a heme propionate, three acidic ligands (Glu-35, Glu-39, and Asp-179), and two water molecules. Using crystallographic methods, this binding site was probed by altering the amount of MnII bound to the protein. Crystals grown in the absence of MnII, or in the presence of EDTA, exhibited diminished electron density at this site. Crystals grown in excess MnII exhibited increased electron density at the proposed binding site but nowhere else in the protein. This suggests that there is only one major MnII binding site in MnP. Crystal structures of a single mutant (D179N) and a double mutant (E35Q,D179N) at this site were determined. The mutant structures lack a cation at the MnII binding site. The structure of the MnII binding site is altered significantly in both mutants, resulting in increased access to the solvent and substrate.
Organizational Affiliation:
Department of Molecular Biology & Biochemistry and Physiology & Biophysics, University of California, Irvine, California 92697-3900, USA.