Design of non-nucleoside inhibitors of HIV-1 reverse transcriptase with improved drug resistance properties. 1.
Hopkins, A.L., Ren, J., Milton, J., Hazen, R.J., Chan, J.H., Stuart, D.I., Stammers, D.K.(2004) J Med Chem 47: 5912-5922
- PubMed: 15537346
- DOI: https://doi.org/10.1021/jm040071z
- Primary Citation of Related Structures:
1TKT, 1TKZ, 1TL1, 1TL3 - PubMed Abstract:
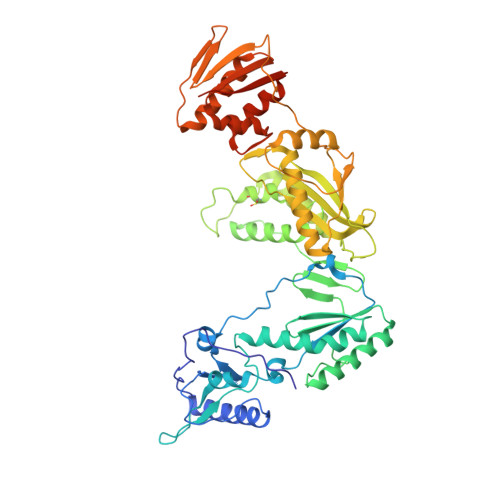
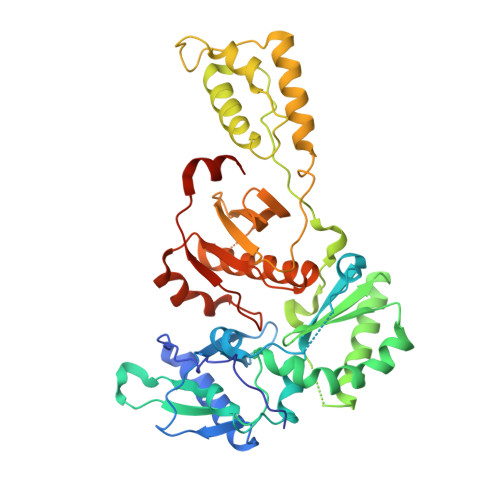
We have used a structure-based approach to design a novel series of non-nucleoside inhibitors of HIV-1 RT (NNRTIs). Detailed analysis of a wide range of crystal structures of HIV-1 RT-NNRTI complexes together with data on drug resistance mutations has identified factors important for tight binding of inhibitors and resilience to mutations. Using this approach we have designed and synthesized a novel series of quinolone NNRTIs. Crystal structure analysis of four of these compounds in complexes with HIV-1 RT confirms the predicted binding modes. Members of this quinolone series retain high activity against the important resistance mutations in RT at Tyr181Cys and Leu100Ile.
Organizational Affiliation:
Division of Structural Biology, The Wellcome Trust Centre for Human Genetics, University of Oxford, Roosevelt Drive, Oxford, OX3 7BN, UK.