Structural determinants of herpesvirus entry mediator recognition by murine B and T lymphocyte attenuator.
Nelson, C.A., Fremont, M.D., Sedy, J.R., Norris, P.S., Ware, C.F., Murphy, K.M., Fremont, D.H.(2008) J Immunol 180: 940-947
- PubMed: 18178834
- DOI: https://doi.org/10.4049/jimmunol.180.2.940
- Primary Citation of Related Structures:
1XAU - PubMed Abstract:
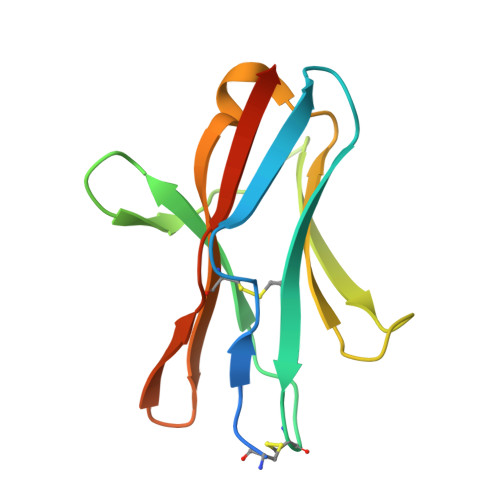
The B and T lymphocyte attenuator (BTLA) appears to act as a negative regulator of T cell activation and growth. BTLA specifically interacts with herpesvirus entry mediator (HVEM), a member of the TNFR family. Herein, we have undertaken surface plasmon resonance studies to quantitatively assess BTLA and HVEM ectodomain interactions. We find that soluble BALB/cJ BTLA engages HVEM with an equilibrium affinity of 0.97+/-0.19 microM while the C57BL/6 BTLA binds slightly better with an equilibrium affinity of 0.42+/-0.06 microM. Despite its lower affinity for HVEM, the kinetic half-life of BALB/cJ BTLA complexes are twice as long as observed for C57BL/6 BTLA (4 vs 2 s). To further explore these interactions, we solved the crystal structure of a murine BTLA (BALB/cJ) ectodomain at 1.8-A resolution, revealing a beta sandwich fold with strong similarity to I-set members of the Ig superfamily. Using a structure-based mutagenesis strategy, we then examined the individual contributions of 26 BTLA surface-exposed residues toward HVEM binding. Four single-site substitutions were identified that decrease HVEM binding below detectable levels and two that decrease binding by more than half. All six of these cluster at the edge of the beta sandwich in a membrane distal patch formed primarily from the A and G strands. This patch falls within the contacting surface recently revealed in the crystal structure of the human BTLA-HVEM cocomplex. The critical binding residues identified here are highly conserved across species, suggesting that BTLA employs a conserved binding mode for HVEM recognition.
Organizational Affiliation:
Department of Pathology and Immunology, Washington University School of Medicine, St. Louis, MO 63110, USA.