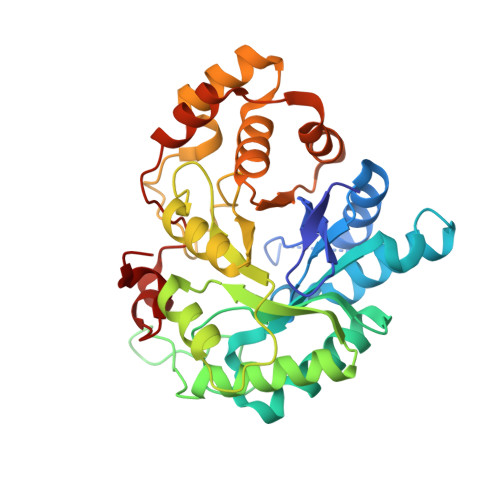
Crystal structures of the multispecific 17beta-hydroxysteroid dehydrogenase type 5: critical androgen regulation in human peripheral tissues
Qiu, W., Zhou, M., Labrie, F., Lin, S.-X.(2004) Mol Endocrinol 18: 1798-1807
- PubMed: 15087468
- DOI: https://doi.org/10.1210/me.2004-0032
- Primary Citation of Related Structures:
1XF0 - PubMed Abstract:
Human type 5 17beta-hydroxysteroid dehydrogenase (17beta-HSD5;AKR1C3) plays a major role in the metabolism of androgens in peripheral tissues. In prostate basal cells, this enzyme is involved in the transformation of dehydroepiandrosterone into dihydrotestosterone, the most potent androgen. It is thus a potential target for prostate cancer therapy because it is understood that the testosterone formation by this enzyme is an important factor, particularly in patients who have undergone surgical or medical castration. Here we report the first structure of a human type 5 17beta-HSD in two ternary complexes, in which we found that the androstenedione molecule has a different binding position from that of testosterone. The two testosterone-binding orientations in the substrate-binding site demonstrate the structural basis of the alternative binding and multispecificity of the enzyme. Phe306 and Trp227 are the key residues involved in ligand recognition as well as product release. A safety belt in the cofactor-binding site enhances nicotinamide adenine dinucleotide phosphate binding and accounts for its high affinity as demonstrated by kinetic studies. These structures have provided a dynamic view of the enzyme reaction converting androstenedione to testosterone as well as valuable information for the development of potent enzyme inhibitors.
Organizational Affiliation:
Oncology and Molecular Endocrinology Research Center, Centre Hospitalier de l'Université Laval Medical Center (CHUL) and Laval University, Sainte-Foy, Quebec, Canada G1V 4G2.