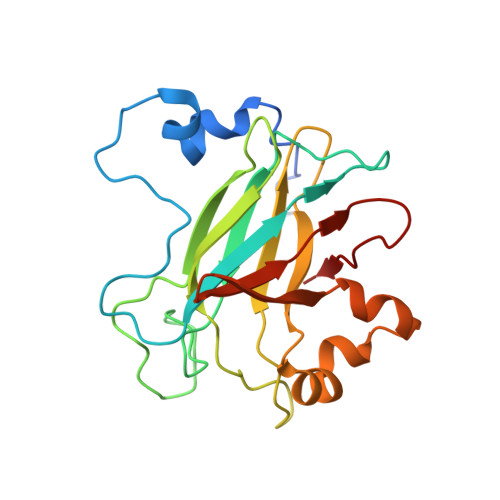
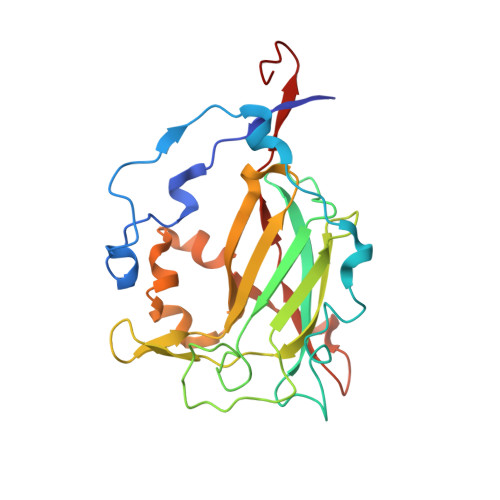
Biophysical Analyses of Designed and Selected Mutants of Protocatechuate 3,4-Dioxygenase
Brown, C.K., Vetting, M.W., Earhart, C.A., Ohlendorf, D.H.(2004) Annu Rev Microbiol 58: 555-585
- PubMed: 15487948
- DOI: https://doi.org/10.1146/annurev.micro.57.030502.090927
- Primary Citation of Related Structures:
2BUM, 2BUQ, 2BUR, 2BUT, 2BUV - PubMed Abstract:
The catechol dioxygenases allow a wide variety of bacteria to use aromatic compounds as carbon sources by catalyzing the key ring-opening step. These enzymes use specifically either catechol or protocatechuate (2,3-dihydroxybenozate) as their substrates; they use a bare metal ion as the sole cofactor. To learn how this family of metalloenzymes functions, a structural analysis of designed and selected mutants of these enzymes has been undertaken. Here we review the results of this analysis on the nonheme ferric iron intradiol dioxygenase protocatechuate 3,4-dioxygenase.
- Center for Metals in Biocatalysis and Department of Biochemistry, Molecular Biology, and Biophysics , Minneapolis, Minnesota 55455, USA. brow0927@umn.edu
Organizational Affiliation: