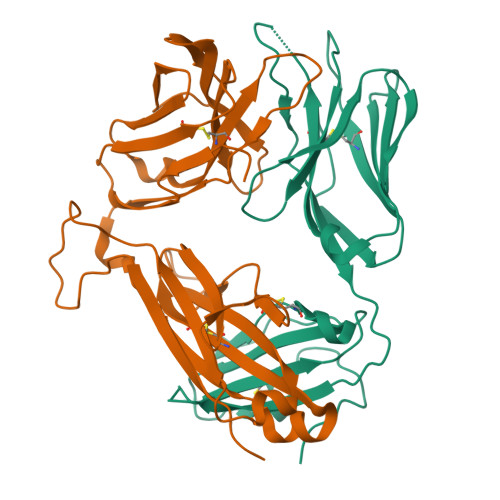
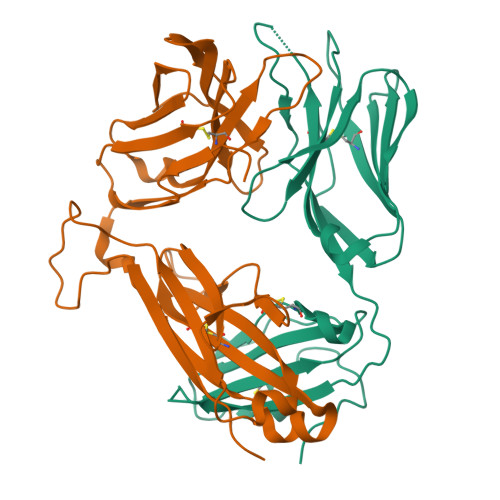
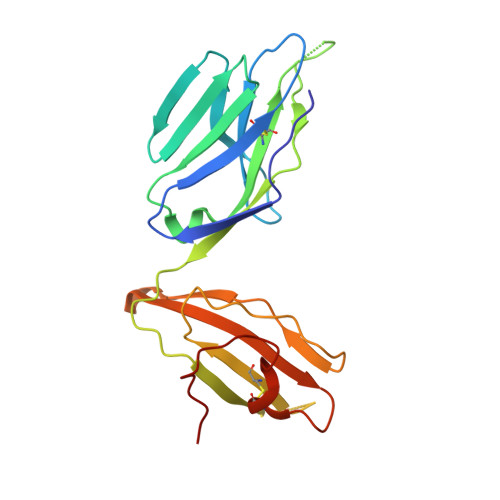
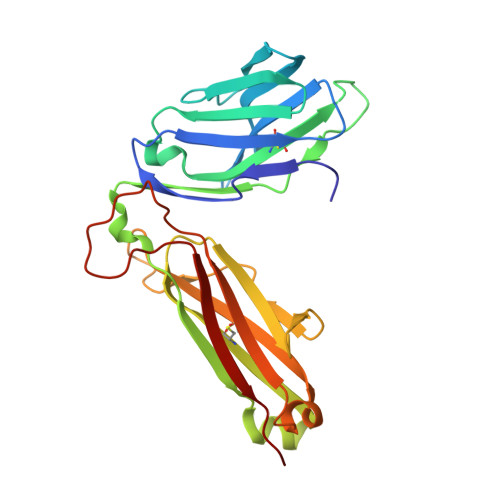
A T cell receptor flattens a bulged antigenic peptide presented by a major histocompatibility complex class I molecule
Tynan, F.E., Reid, H.H., Kjer-Nielsen, L., Miles, J.J., Wilce, M.C., Kostenko, L., Borg, N.A., Williamson, N.A., Beddoe, T., Purcell, A.W., Burrows, S.R., McCluskey, J., Rossjohn, J.(2007) Nat Immunol 8: 268-276
- PubMed: 17259989
- DOI: https://doi.org/10.1038/ni1432
- Primary Citation of Related Structures:
2NW2, 2NW3, 2NX5 - PubMed Abstract:
Plasticity of the T cell receptor (TCR) is a hallmark of major histocompatibility complex (MHC)-restricted T cell recognition. However, it is unclear whether interactions of TCR and peptide-MHC class I (pMHCI) always conform to this paradigm. Here we describe the structure of a TCR, ELS4, in its non-ligand-bound form and in complex with a prominent 'bulged' Epstein-Barr virus peptide bound to HLA-B(*)3501. This complex was atypical of previously characterized TCR-pMHCI interactions in that a rigid face of the TCR crumpled the bulged antigenic determinant. This peptide 'bulldozing' created a more featureless pMHCI determinant, allowing the TCR to maximize MHC class I contacts essential for MHC class I restriction of TCR recognition. Our findings represent a mechanism of antigen recognition whereby the plasticity of the T cell response is dictated mainly by adjustments in the MHC-bound peptide.
- Protein Crystallography Unit, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: