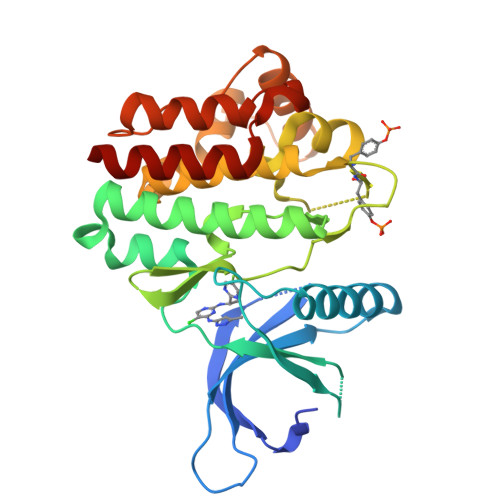
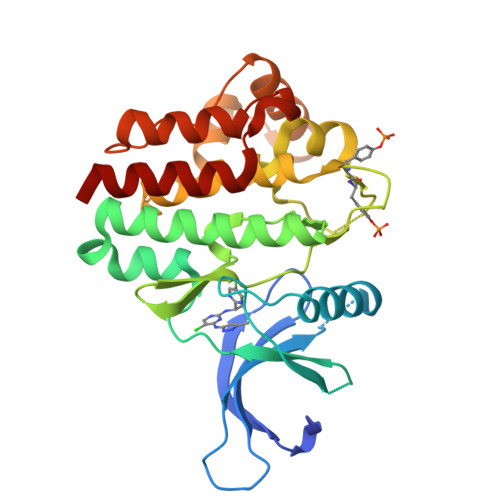
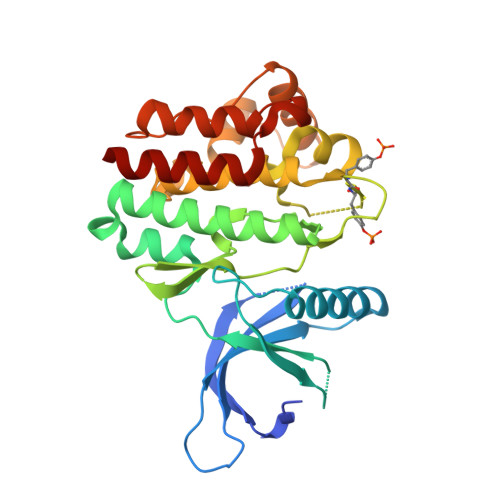
Discovery of 5-Chloro-N2-[(1S)-1-(5-Fluoropyrimidin-2-Yl) Ethyl]-N4-(5-Methyl-1H-Pyrazol-3-Yl)Pyrimidine-2,4-Diamine (Azd1480) as a Novel Inhibitor of the Jak/Stat Pathway
Ioannidis, S., Lamb, M.L., Wang, T., Almeida, L., Block, M.H., Davies, A.M., Peng, B., Su, M., Zhang, H., Hoffmann, E., Rivard, C., Green, I., Howard, T., Pollard, H., Read, J., Alimzhanov, M., Bebernitz, G., Bell, K., Ye, M., Huszar, D., Zinda, M.(2011) J Med Chem 54: 262
- PubMed: 21138246
- DOI: https://doi.org/10.1021/jm1011319
- Primary Citation of Related Structures:
2XA4 - PubMed Abstract:
The myeloproliferative neoplasms, polycythemia vera, essential thrombocythemia, and idiopathic myelofibrosis are a heterogeneous but related group of hematological malignancies characterized by clonal expansion of one or more myeloid lineages. The discovery of the Jak2 V617F gain of function mutation highlighted Jak2 as a potential therapeutic target in the MPNs. Herein, we disclose the discovery of a series of pyrazol-3-yl pyrimidin-4-amines and the identification of 9e (AZD1480) as a potent Jak2 inhibitor. 9e inhibits signaling and proliferation of Jak2 V617F cell lines in vitro, demonstrates in vivo efficacy in a TEL-Jak2 model, has excellent physical properties and preclinical pharmacokinetics, and is currently being evaluated in Phase I clinical trials.
Organizational Affiliation:
Department of Cancer Chemistry, AstraZeneca R&D, Boston, Massachusetts, United States. stephanos.ioannidis@astrazeneca.com