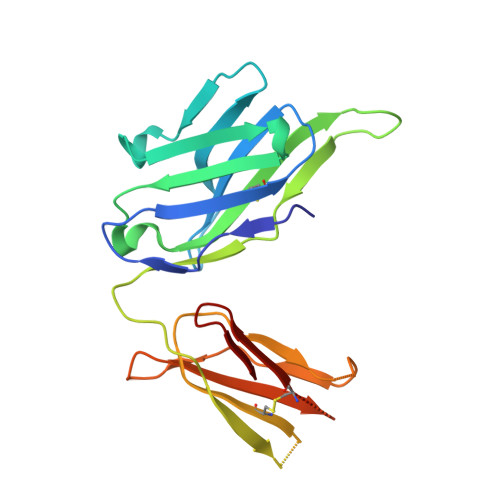
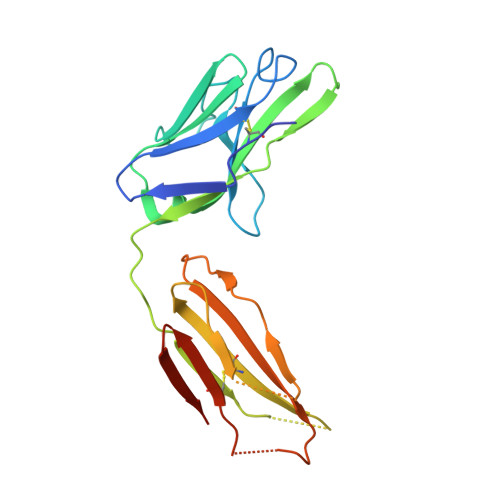
Different Binding Modes of Free and Carrier-Protein-Coupled Nicotine in a Human Monoclonal Antibody.
Tars, K., Kotelovica, S., Lipowsky, G., Bauer, M., Beerli, R., Bachmann, M., Maurer, P.(2012) J Mol Biol 415: 118
- PubMed: 22079050
- DOI: https://doi.org/10.1016/j.jmb.2011.10.042
- Primary Citation of Related Structures:
2YK1, 2YKL - PubMed Abstract:
Nicotine is the principal addictive component of tobacco. Blocking its passage from the lung to the brain with nicotine-specific antibodies is a promising approach for the treatment of smoking addiction. We have determined the crystal structure of nicotine bound to the Fab fragment of a fully human monoclonal antibody (mAb) at 1.85 Å resolution. Nicotine is almost completely (>99%) buried in the interface between the variable domains of heavy and light chains. The high affinity of the mAb is the result of a charge-charge interaction, a hydrogen bond, and several hydrophobic contacts. Additionally, similarly to nicotinic acetylcholine receptors in the brain, two cation-π interactions are present between the pyrrolidine charge and nearby aromatic side chains. The selectivity of the mAb for nicotine versus cotinine, which is the major metabolite of nicotine and differs in only one oxygen atom, is caused by steric constraints in the binding site. The mAb was isolated from B cells of an individual immunized with a nicotine-carrier protein conjugate vaccine. Surprisingly, the nicotine was bound to the Fab fragment in an orientation that was not compatible with binding to the nicotine-carrier protein conjugate. The structure of the Fab fragment in complex with the nicotine-linker derivative that was used for the production of the conjugate vaccine revealed a similar position of the pyridine ring of the nicotine moiety, but the pyrrolidine ring was rotated by about 180°. This allowed the linker part to reach to the Fab surface while high-affinity interactions with the nicotine moiety were maintained.
Organizational Affiliation:
Biomedical Research and Study Center, Ratsupites 1, Riga LV 1067, Latvia. kaspars@biomed.lu.lv