Filling a Hole in Cytochrome P450 Bm3 Improves Substrate Binding and Catalytic Efficiency.
Huang, W.-C., Westlake, A.C.G., Marechal, J.-D., Joyce, M.G., Moody, P.C.E., Roberts, G.C.K.(2007) J Mol Biol 373: 633
- PubMed: 17868686
- DOI: https://doi.org/10.1016/j.jmb.2007.08.015
- Primary Citation of Related Structures:
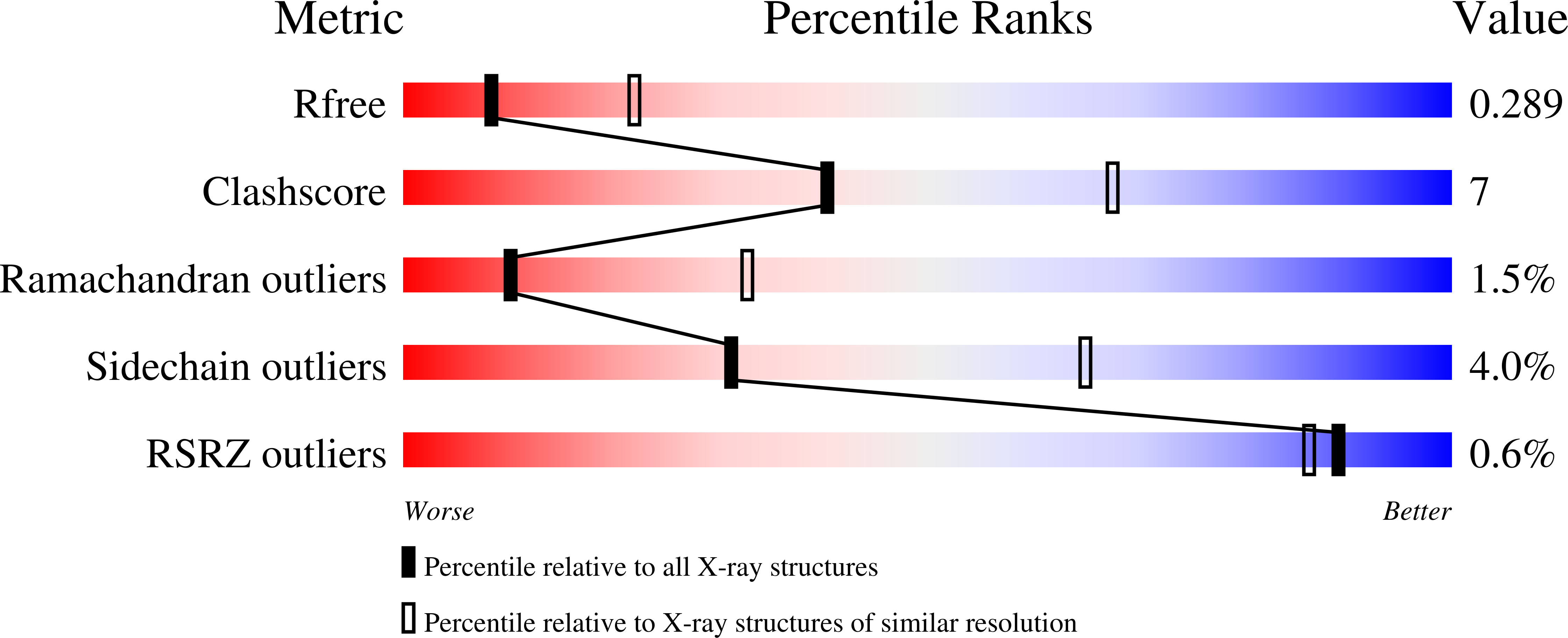
2UWH - PubMed Abstract:
Cytochrome P450BM3 (CYP102A1) from Bacillus megaterium, a fatty acid hydroxylase, is a member of a very large superfamily of monooxygenase enzymes. The available crystal structures of the enzyme show non-productive binding of substrates with their omega-end distant from the iron in a hydrophobic pocket at one side of the active site. We have constructed and characterised mutants in which this pocket is filled by large hydrophobic side-chains replacing alanine at position 82. The mutants having phenylalanine or tryptophan at this position have very much (approximately 800-fold) greater affinity for substrate, with a greater conversion of the haem iron to the high-spin state, and similarly increased catalytic efficiency. The enzyme as isolated contains bound palmitate, reflecting this much higher affinity. We have determined the crystal structure of the haem domain of the Ala82Phe mutant with bound palmitate; this shows that the substrate is binding differently from the wild-type enzyme but still distant from the haem iron. Detailed analysis of the structure indicates that the tighter binding in the mutant reflects a shift in the conformational equilibrium of the substrate-free enzyme towards the conformation seen in the substrate complex rather than differences in the enzyme-substrate interactions. On this basis, we outline a sequence of events for the initial stages of the catalytic cycle. The Ala82Phe and Ala82Trp mutants are also very much more effective catalysts of indole hydroxylation than the wild-type enzyme, suggesting that they will be valuable starting points for the design of mutants to catalyse synthetically useful hydroxylation reactions.
Organizational Affiliation:
Henry Wellcome Laboratories of Structural Biology, Department of Biochemistry, University of Leicester, Leicester LE1 9HN, UK.